Do you (want to) use whole-plasmid sequencing (e.g. @plasmidsaurus) but are put off by $15/sample? Use SAVEMONEY—our (postdoc Masaaki Uematsu’s) algorithm that lets you mix 6 or more plasmids/sample & computationally de-mixes reads!
https://colab.research.google.com/github/MasaakiU/MultiplexNanopore/blob/master/colab/MultiplexNanopore.ipynb
https://www.biorxiv.org/content/10.1101/2023.04.12.536413v1
How it works: 1/n
Multiplexing exists for long-read sequencing but typically involves extra barcoding or is geared toward those with their own sequencers. SAVEMONEY (Simple Algorithm for Very Efficient Multiplexing of Oxford Nanopore Experiments for You) is for users of 3rd party services. 2/n
Nanopore sequencing typically provides 100s of reads/sample to generate a consensus sequence. But Masaaki realized that w/prior information of expected sequence & Bayesian analysis, you can get sequences w/~30 reads/plasmid—in other words, money is being left on the table. 3/n
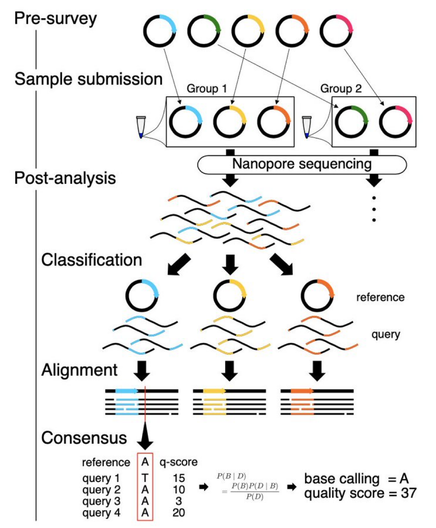
SAVEMONEY has two basic parts: (1) pre-survey & (2) post-analysis. Pre-survey examines your plasmid maps and tells you which groups of plasmids to mix together. Post-analysis classifies, aligns, and generates consensus sequences for each plasmid in the mixture. 4/n
Incredibly, you can reliably mix together plasmids differing by as few as 2 bases & up to 6 (or even more) plasmids in a single sample, which lowers the effective cost of whole-plasmid sequencing to less than that of a single Sanger run! We hope that SAVEMONEY will... 5/n