Evelien Adriaenssens
- 150 Followers
- 75 Following
- 25 Posts
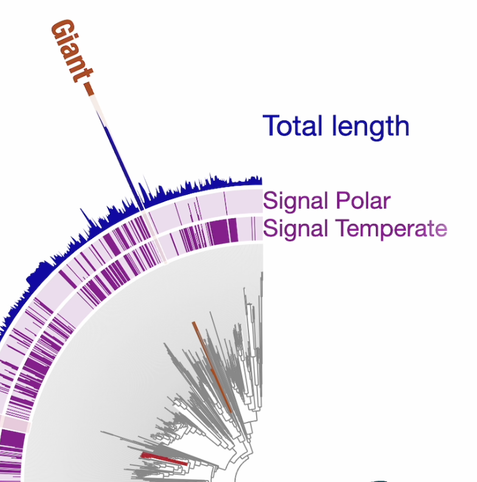
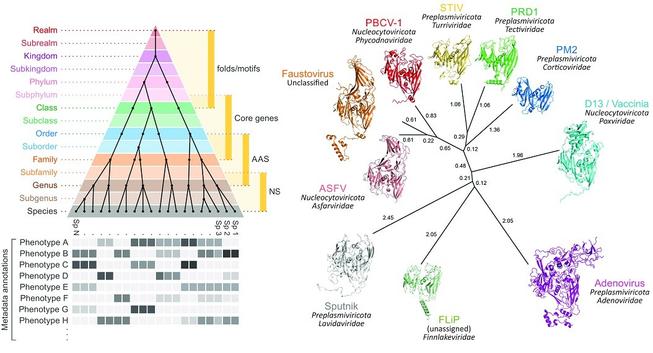
Four principles to establish a universal virus taxonomy
Consensus view to reconcile traditional ways of virus classification with a phylogenetic-based one that can accommodate modern virus discovery approaches
Virology friends, read on 👇🏼👇🏼
@PLOSBiology
---
RT @PLOSBiology
#Viral #taxonomy: transforming the existing classification of #viruses into a phylogenetic one is an ongoing challenge. This Consensus View explains how such #ta…
https://twitter.com/PLOSBiology/status/1625438878411771904
PLOS Biology on Twitter
“#Viral #taxonomy: transforming the existing classification of #viruses into a phylogenetic one is an ongoing challenge. This Consensus View explains how such #taxonomy can encapsulate viral diversity & recognize independent biological origins #PLOSBiology https://t.co/VCBiM0udaU”
Definitely a team effort to get this one out. :)
We've isolated a novel phage that can disrupt established biofilms ... but it doesn't encode a depolymerase or exhibit depolymerase activity. I think this one is going to keep us busy for a long time.
Phage vB_KmiS-Kmi2C infects members of the Klebsiella oxytoca complex and represents a novel genus of lytic bacteriophage https://www.biorxiv.org/content/10.1101/2023.01.12.523727v1
#Klebsiella #phage #biofilms #Klebsiellaoxytoca #Klebsiellamichiganensis
Molecular mechanisms of antibiotic resistance revisited - Nature Reviews Microbiology
In this Review, Blair, Webber and colleagues explore our understanding of the mechanisms of antibiotic resistance, including reduced permeability, antibiotic efflux, modification or alteration of the antibiotic target, modification or destruction of the drug itself, and bypass of metabolic pathways. They also discuss how this information can aid in developing the next generation of antimicrobial therapies.
Good luck!