I could make the night train journey there thanks to @Co_Biologists's DMM Conference Travel Grant - much appreciated!
#lightsheetmicroscopy #tissueclearing #rodent #neuroanatomy #neuroscience
I could make the night train journey there thanks to @Co_Biologists's DMM Conference Travel Grant - much appreciated!
🔬📄 'Efficacy and Feasibility of Tissue-Clearing Technique and Three-Dimensional Imaging in the Human Gastrointestinal Tissues Using Illuminate Cleared Organs to Identify Target Molecules' - a Karger: #Gastroenterology article on #ScienceOpen -
🔗 https://www.scienceopen.com/document?vid=09d6ca4d-9131-4734-89df-86fae09bc8d7
#TissueClearing #3DImaging #LUCIDProtocol #Pathology #Microscopy
Efficacy and Feasibility of Tissue-Clearing Technique and Three-Dimensional Imaging in the Human Gastrointestinal Tissues Using Illuminate Cleared Organs to Identify Target Molecules
<p xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance" dir="auto" id="d7337466e239"> <b> <i>Introduction:</i> </b> Tissue-clearing technology has shown potential for comprehensive structural and functional analysis through three-dimensional (3D) imaging of biological tissue. However, its effectiveness in human specimens remains insufficiently explored. In this study, we validated the illuminate cleared organs to identify target molecules (LUCID) protocol for human gastrointestinal specimens and demonstrated its utility in enhancing tissue transparency and 3D imaging. <b> <i>Methods:</i> </b> The gastrointestinal mucosa specimens resected via endoscopic submucosal dissection including the esophagus, stomach, duodenum, and colon were fluorescently stained and optically cleared using LUCID. Cleared specimens were imaged in 3D form by confocal laser scanning microscope, and the observable depth at any five points was measured and compared to non-cleared specimens, respectively. After clearing and imaging, the specimens were restored to the formalin-fixed paraffin-embedded form again, and conventional two-dimensional pathological evaluation using hematoxylin-eosin, Ki67, p53, and E-cadherin staining was performed to compare them with their preclearing state. <b> <i>Results:</i> </b> The observable depth was significantly extended after clearing for specimens from each organ (esophagus 228.3 ± 14.9 µm vs. 1,036.7 ± 62.9 µm, <i>p</i> < 0.05; stomach 115.2 ± 5.5 µm vs. 428.7 ± 15.9 µm, <i>p</i> < 0.05; duodenum 256.2 ± 9.5 µm vs. 787.0 ± 18.6 µm, <i>p</i> < 0.05, colon 113.9 ± 5.4 µm vs. 436.6 ± 18.5 µm, <i>p</i> < 0.05). The pathological evaluation after clearing revealed a preserved fine structure and staining and showed no apparent deformation, degeneration, or tissue damage compared with before clearing. <b> <i>Conclusions:</i> </b> The effectiveness of tissue clearing using LUCID on human gastrointestinal specimens was demonstrated, and the LUCID protocol had minimal impact on specimen morphology and staining. LUCID is expected to be a method that enables comprehensive structural analysis of human gastrointestinal mucosa and lesions that may avoid missing microscopic findings that may occur in split-face pathological assessment. </p>
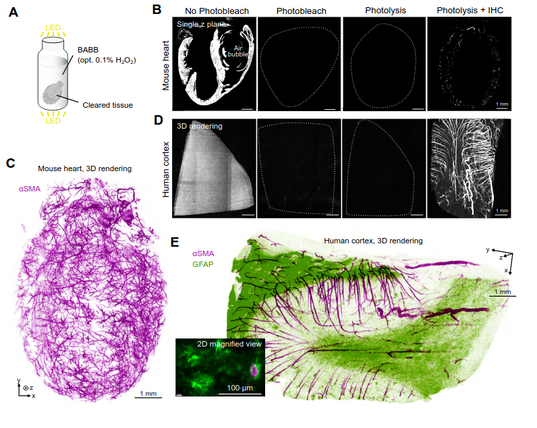
Here's an interesting way to do photobleaching with cleared tissue: Add 0.1% H2O2 to the clearing medium and blast with LEDs for 24h at RT. Also includes very detailed instructions for building an LED photobleaching setup - the samples ate placed between two PCBs with LEDs in a ventilated box.
Open-source Photobleacher for Fluorescent Imaging of Large Pigment-Rich Tissues
Murakami et al., preprint at biorxiv 2025
https://www.biorxiv.org/content/10.1101/2025.02.24.639965v2.full.pdf
New publication from Kwanghun Chung lab at MIT improves on their SWITCH approach and eFLASH stochastic electrotransport to improve antibody labeling by gradually shifting the microenvironment. Looks interesting, I wonder how well it can be adapted to other labeling protocols.
Uniform volumetric single-cell processing for organ-scale molecular phenotyping
Yun et al., Nature Biotechnology 2025
https://doi.org/10.1038/s41587-024-02533-4
#neuroscience #tissueclearing #lightsheet #microscopy #fluorescenceFriday
Wow, TRISCO (née TRIC-DISCO) is out in Science! This cool approach allows imaging of mRNA transcripts throughout the cleared mouse brain. It uses a signal amplification step with in situ Hybridization Chain Reaction (isHCR) and combines it with DISCO-style solvent-based #tissueclearing to make the brain transparent.
Whole-brain spatial transcriptional analysis at cellular resolution
Kanatani et al., Science 2024
https://doi.org/10.1126/science.adn9947
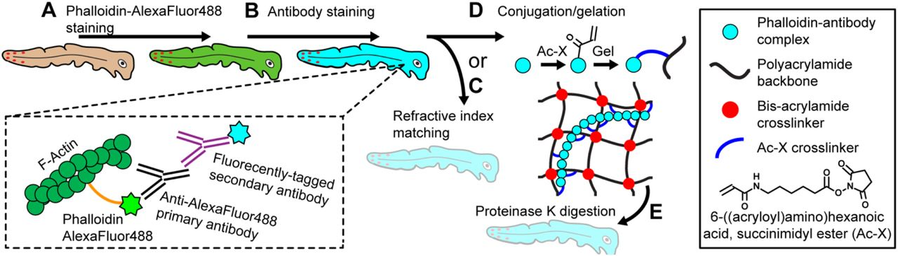
Clever stacking of labels allows the retention of cytoskeleton in tissues (here: Shark skin):
Phalloidin-Alexa488 -> Anti-Alexa488 primary antibody -> Secondary antibody -> Hydrogel crosslinking -> ProtK tissue digestion -> Clearing
Whole Tissue Imaging of Cellular Boundaries at Sub-Micron Resolutions for Automatic Cell Segmentation: Applications in Epithelial Bending of Ectodermal Appendages
Norris et al., preprint at biorxiv 2024
https://doi.org/10.1101/2024.06.26.600880
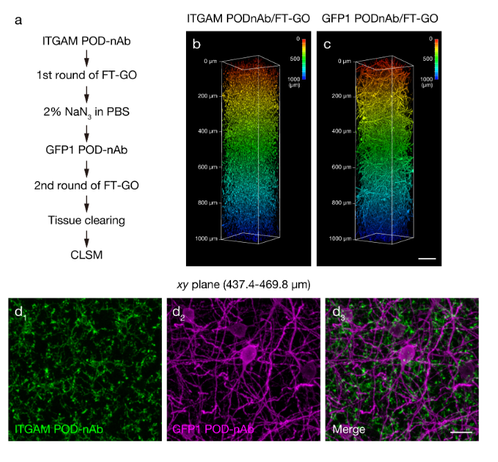
Interesting, though complex, signal amplification strategy using Nanobodies coupled to peroxidase -> added Glucose Peroxidase -> (reaction glucose -> H2O2) -> tyramide-based signal amplification -> fluorophore deposition.
A Three Dimensional Immunolabeling Method with Peroxidase-fused Nanobodies and Fluorochromized Tyramide-Glucose Oxidase Signal Amplification
Yamauchi et al., preprint at biorxiv 2024
https://doi.org/10.1101/2024.10.25.620157
In case anyone was following the recent Science paper about using Tartrazine for #tissueclearing, apparently there is at least one (competing) group that couldn't reproduce it and wrote a preprint about it:
Tartrazine cannot make live tissues transparent
https://doi.org/10.1101/2024.09.29.615648
Discussion on Pubpeer, including the Tartrazine paper's author's response:
https://pubpeer.com/publications/81314BB3706B3D6ADAD49F301B2FA5
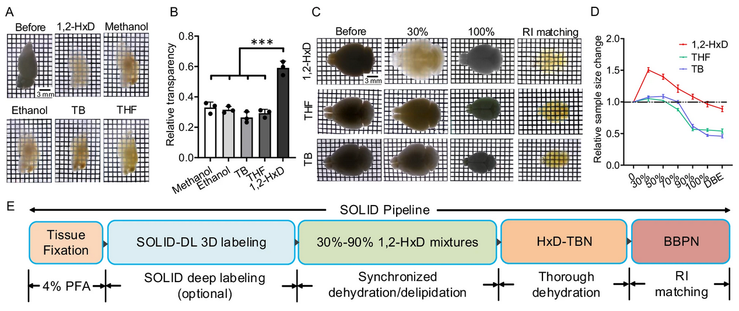
New solvent-based #tissueclearing protocol claims reduced tissue distortion: Dehydration/delipidation with Hexanediol, tert-Butanol and N-butyldiethanolamine, and clearing with Benzyl Benzoate / PEGMA / N-butyldiethanolamine.
SOLID: minimizing tissue distortion for brain-wide profiling of diverse architectures
Zhu et al., Nature Comms 2024
https://doi.org/10.1038/s41467-024-52560-7