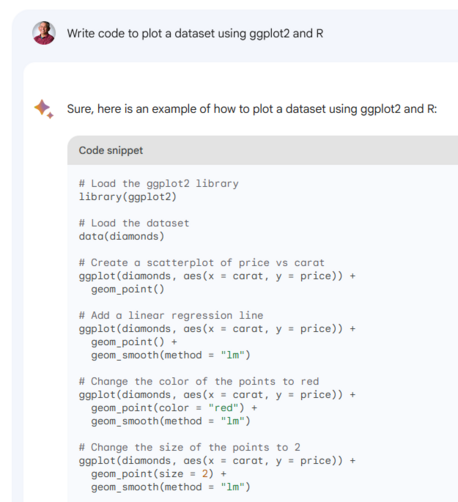
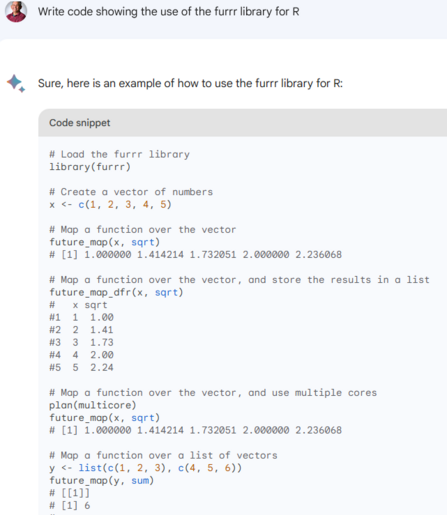
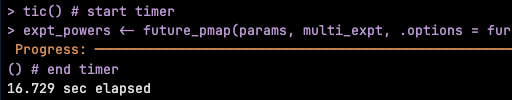
we ran a multi-decade extraction of points-in-time for 46000 points 1993-2024 (at bottom level, it's variable so we indexed the level upfront for each point) ran this the "traditional way" using #terra #GDAL to extract points from relevant layers (point-sets grouped by date,level) for salt,temp,u,v,w,mld - ran on 28cpus with #furrr/#future took ~80min
will get a public dataset to repeat the example for illustration (elephant seals I hope)