🧬 Is it possible to make whole-genome sequencing practical for everyday clinical care?
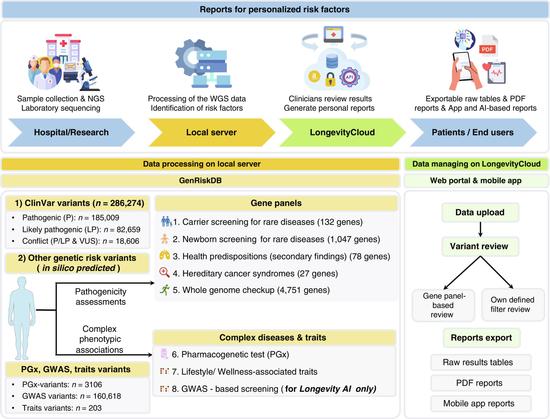
🔗 GenRiskPro: A Comprehensive Whole-Genome Sequencing Analysis Platform for Clinical and Wellness Applications. https://spj.science.org/doi/epdf/10.34133/csbj.0011
📚 CSBJ (Smart Hospital Section) - A Science Partner Journal: https://spj.science.org/page/csbj/for-authors#smarthospital
#Genomics #PrecisionMedicine #WholeGenomeSequencing #DigitalHealth #PersonalizedMedicine