It’s really interesting to me how similar ONT and Apple are with their presentations and UX 🍎
@nanopore My selected novel statements from ONT's Tech Update from the 2024 Nanopore Community meeting:
* Updated documentation centre is public: no need to log in
* For large-scale human sequencing (thousands and thousands of individuals), >90% runs over 100 Gb (30X)
* 24-plex direct RNA barcoding kit
* Improved enzyme motor speed, ONT have got a flow cell over 270 Gb yield
@nanopore Should probably get this link out sometime before the conference ends, just in case it interests anyone...
My notes for London Calling 2024:
Details are available here: https://doi.org/10.1515/jib-2023-0033
#NanoporeConf #Genomics @PuckerLab
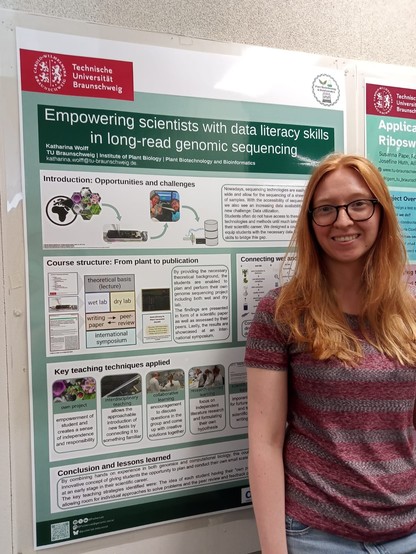
Data literacy in genome research
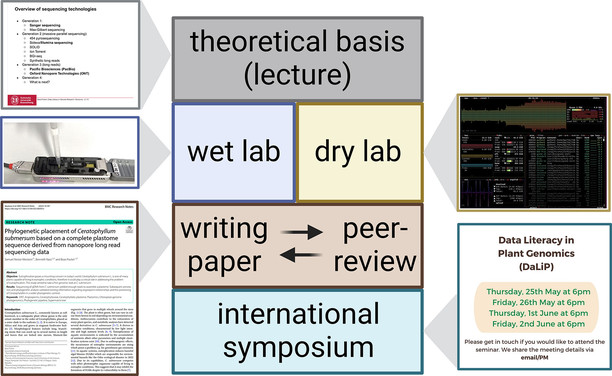
With an ever increasing amount of research data available, it becomes constantly more important to possess data literacy skills to benefit from this valuable resource. An integrative course was developed to teach students the fundamentals of data literacy through an engaging genome sequencing project. Each cohort of students performed planning of the experiment, DNA extraction, nanopore sequencing, genome sequence assembly, prediction of genes in the assembled sequence, and assignment of functional annotation terms to predicted genes. Students learned how to communicate science through writing a protocol in the form of a scientific paper, providing comments during a peer-review process, and presenting their findings as part of an international symposium. Many students enjoyed the opportunity to own a project and to work towards a meaningful objective.
https://doi.org/10.1515/jib-2023-0033
#nanoporeconf #Genomics #londoncalling @PuckerLab
Data literacy in genome research
With an ever increasing amount of research data available, it becomes constantly more important to possess data literacy skills to benefit from this valuable resource. An integrative course was developed to teach students the fundamentals of data literacy through an engaging genome sequencing project. Each cohort of students performed planning of the experiment, DNA extraction, nanopore sequencing, genome sequence assembly, prediction of genes in the assembled sequence, and assignment of functional annotation terms to predicted genes. Students learned how to communicate science through writing a protocol in the form of a scientific paper, providing comments during a peer-review process, and presenting their findings as part of an international symposium. Many students enjoyed the opportunity to own a project and to work towards a meaningful objective.
https://doi.org/10.1101/2024.02.14.580303
#nanoporeconf #Genomics #londoncalling @PuckerLab
#NanoporeConf @[email protected] I think @[email protected] is selling themselves short with regards to number of genomes per PromethION flow cell. With 15kb reads, the coverage required to hit every base is about 10X.
With 650bp reads, the required coverage is about 15X.
What I'm saying is that because of longer reads, ONT doesn't need to do 40X for a human genome. They're already at 3 genomes per flow cell with 150 Gb output.
https://www.reddit.com/r/genomics/comments/z2sj1q/comment/ixj1rgg/?context=3
#NanoporeConf @joepdl started his talk on targeted sequencing of infectious diseases with a mihi, then a content warning for COVID, then a "results may differ depending on your government" disclaimer.
Great to see a little bit of fatalistic humour to brighten my day!