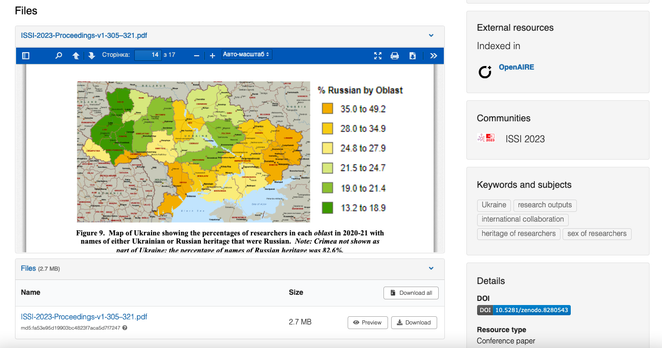
Strange methodology: "We noted that only Ukrainians had surnames ending in chuk, iuk or skyi, and only Russians had surnames ending in (e.g., Lenin, Putin, Stalin). These lists of names with definite heritage enabled us to mark many of the names of the researchers in each oblast as either Russian (RU) or Ukrainian (UA)."
👉 https://doi.org/10.5281/zenodo.8280543
For decades, the Soviets forced Ukrainians to change their surnames to Russian. Excuse me, but these #ISSI2023 paper reek of Kremlin narratives. 🤬
Researchers in Ukraine: output, international collaboration, impact, heritage and sex, 1995 to 2021
The invasion of Ukraine by the Russian army in February 2022 caused us to look back at Ukrainian science before the war and appraise its strengths and weaknesses, so that after peace and reconstruction it may flourish again and be better attuned to the new era. We examined its publications recorded in the Web of Science (WoS) from 1995 to 2021, and evaluated their quantity and impact, their major fields and Ukraine’s international partners. We also looked at the outputs of the 26 individual oblasts or regions, and the heritage of the researchers in each, Ukrainian or Russian, based on their surnames and given names. Ukrainian scientific output, at about 0.5% of that of the world, was high relative to its wealth, but its impact was rather low, probably because of a lack of contestable funding. It was skewed towards the physical sciences and away from biology and medicine, and so may have led to poorer health in Ukraine than in its geographical neighbours. Its researchers were more likely to be of Ukrainian than Russian heritage the further west they were located. Women were well represented in Ukrainian science and formed more than half the total in most oblasts.