Identification of FtsZ Protein Nucleotide-Binding Site Effectors Based on Cheminformatics and Structural-Biological Analysis - Cytology and Genetics
There is a large group of bacterial FtsZ inhibitors whose biological activity has been confirmed biochemically. However, the sites of protein–ligand interaction for most of them remain unknown, significantly complicating the further search and combinatorial design of FtsZ inhibitors. This study presents the results of bioinformatic analysis of bacterial FtsZ effectors, the interaction of which has been proven and documented in the ChEMBL database of biologically active molecules. With an integrated approach based on chemo- and bioinformatic methods and AI-based predictions, 23 inhibitors of nucleotide-binding site (NBS), as well as 16 new effectors of the interdomain cleft (IDC), were identified.

Identification of FtsZ Interdomain Cleft Effectors Based on Pharmacophore Search and Molecular Docking - Cytology and Genetics
Abstract There are a significant number of inhibitors of the bacterial FtsZ protein with biochemically confirmed biological activity, at the same time, their binding sites remain unclear. This significantly complicates further combinatorial design, and, in the current study, the results of a computational search for effectors of the Inter-Domain Cleft (IDC) site are presented. The actual research was based on the results of pharmacophore screening using the Pharmit service and molecular docking with CCDC GOLD and iGEMDOCK programs. The objective group was a combined library of 379 compounds, which was designed based on revision of the structural database of the RCSB Protein Data Bank and compounds from the ChEMBL database, for which direct interaction with FtsZ has been proven biochemically. According to the results of pharmacophore search, docking, and structural analysis, 88 effectors of the IDC site were identified. One more curcumin compound has been identified as a potential IDC site effector.
Membrane trafficking pathways play a prominent role in immunity of plants against diseases.
#Effectors of
#plant #pathogens have evolved to interfere with aspects of membrane transport systems and reprogram host plant vesicle traffic to overcome plant
#resistance. Review by Enoch Lok Him Yuen and others -
https://doi.org/10.1146/annurev-phyto-021622-123232Very happy to share our preprint on fishing for
#plant targets of ADP-ribosylating bacterial type III
#effectors - great team effort, 🙏 thanks to everyone involved! (finally incl. some halfway decent MS2 spectra of the modification, 9 years later...😉)
#slowscience #plantscience #ADPribosylation #plantimmunity https://biologists.social/@biorxivpreprint/111132762774432947Untargeted proteomics identifies plant substrates of the bacterial-derived ADP-ribosyltransferase AvrRpm1 https://www.biorxiv.org/content/10.1101/2023.09.25.558804v1?med=mas
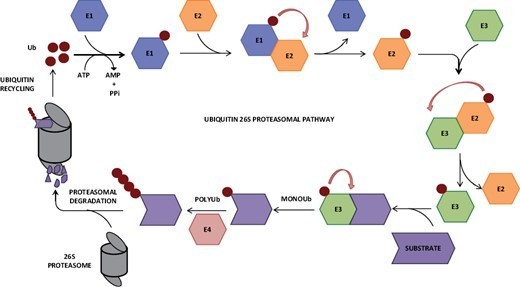
This #review summarises how pathogenic #effectors can mimic or target the highly conserved #ubiquitin–#proteasome system to disrupt cellular processes in their #plant hosts.
https://doi.org/10.1093/jxb/erad191
#ubiquitination #immunity #PAMP #effector

Ubiquitination from the perspective of plant pathogens
A review focusing on the strategies adopted by plant pathogens to secrete effectors that either mimic E3 ligase properties or target the ubiquitin–proteasome sy
#Effectors play a central role in determining the outcome of
#plant−
#pathogen interactions and can provide fundamental knowledge of microbial pathogenesis and disease resistance. Review of plant pathogen effectors and the recently developed analytic tools and technologies for identifying them - article in Phytopathology (2023, vol. 113, issue 4, pp. 637-650) by Amelia H. Lovelace and others like
@michhulin -
https://doi.org/10.1094/PHYTO-09-22-0337-KD 

Effector Identification in Plant Pathogens | Phytopathology®
Effectors play a central role in determining the outcome of plant−pathogen interactions. As key virulence proteins, effectors are collectively indispensable for disease development. By understanding the virulence mechanisms of effectors, fundamental knowledge of microbial pathogenesis and disease resistance have been revealed. Effectors are also considered double-edged swords because some of them activate immunity in disease resistant plants after being recognized by specific immune receptors, which evolved to monitor pathogen presence or activity. Characterization of effector recognition by their cognate immune receptors and the downstream immune signaling pathways is instrumental in implementing resistance. Over the past decades, substantial research effort has focused on effector biology, especially concerning their interactions with virulence targets or immune receptors in plant cells. A foundation of this research is robust identification of the effector repertoire from a given pathogen, which depends heavily on bioinformatic prediction. In this review, we summarize methodologies that have been used for effector mining in various microbial pathogens which use different effector delivery mechanisms. We also discuss current limitations and provide perspectives on how recently developed analytic tools and technologies may facilitate effector identification and hence generation of a more complete vision of host−pathogen interactions. Copyright © 2023 The Author(s). This is an open access article distributed under the CC BY-NC-ND 4.0 International license.
#High-Throughput Discovery and Characterization of #Viral Transcriptional #Effectors in Human Cells | bioRxiv
https://www.biorxiv.org/content/10.1101/2022.12.16.520835v1
#synbio