🧬 Why are we still studying genes in isolation when biology works as a network?
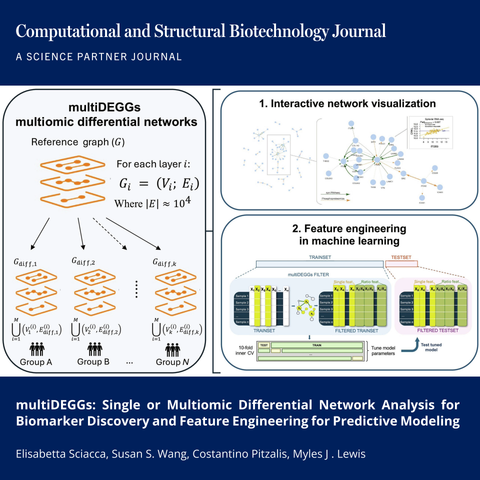
🔗 multiDEGGs: Single or Multiomic Differential Network Analysis for Biomarker Discovery and Feature Engineering for Predictive Modeling. Computational and Structural Biotechnology Journal (CSBJ). DOI: https://doi.org/10.34133/csbj.0001
📚 CSBJ - A Science Partner Journal: https://spj.science.org/journal/csbj
#Bioinformatics #Multiomics #MachineLearning #PrecisionMedicine #DataScience #Biomarkers