RT @theChaudhurilab
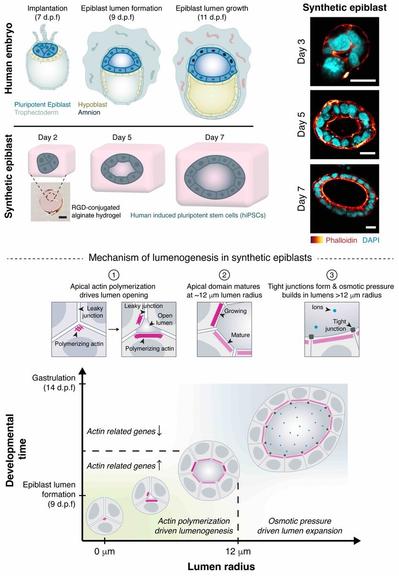
New preprint alert! 🚨 Excited to share @DhirajIndana’s phenomenal work on the mechanism of lumen formation in a human epiblast model. A short thread 🧵
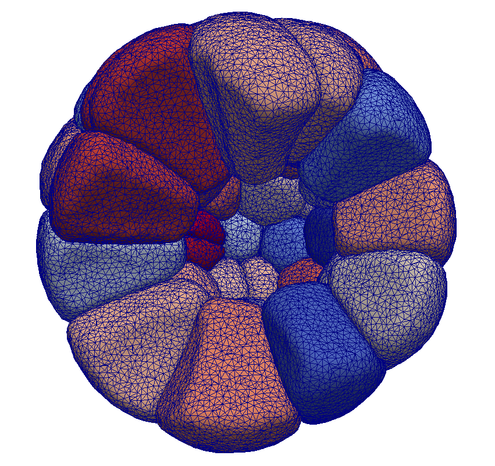
https://www.biorxiv.org/content/10.1101/2023.04.20.537711v1Congratulations to Roman Vetter from our group for winning an award in the @snf_ch #image #competition with one of his stunning videos of tissue growth:
https://www.flickr.com/photos/snsf_image_competition/52797439078/in/album-72177720307288892/
#SwissScienceVideo
Stay tuned for his paper!!

"Tumour growth simulation" by Roman Vetter, ETH Zurich
RT @EbisuyaMiki
W/ the spirit of St Jordi🐉, I read my favorite book (only 3 chaps, though).
"Scaling: Why is Animal Size so Important?" taught me animal size dictates many parameters, including time. Rather than real clock time, using physiological time helps understand animals comprehensively
RT @ClotildeCadart
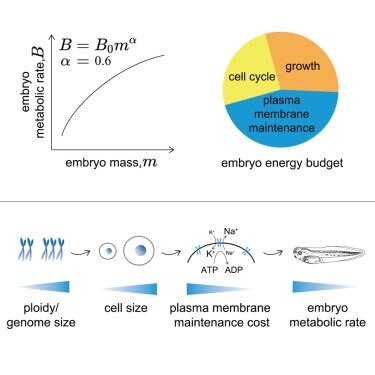
Our paper is out! Xenopus embryos follow
#kleiber’s power law between metabolic rate and body mass. But triploids embryos which are made of fewer larger cells have a lower metabolic rate than diploids. 1/3

Dagmar Iber on Twitter
“Excited to introduce #SimuCell3D, an #opensource 3D cell-based model to simulate large and complex tissues at high resolution.
Check out our latest @biorxivpreprint https://t.co/ZA13Wsvd0g
A short summary thread about the software tool:🧵(1/9)”
RT @virtual_embryo
Can we predict cellular forces just from microscopy ?
We are happy to release a robust method by @SachaIchbiah to create 3D atlases of relative cell mechanics in embryos or tissues from fluorescent images of cell outlines:
https://tinyurl.com/5fj2c4b8
A thread (1/n)
Excited to introduce #SimuCell3D, an #opensource 3D cell-based model to simulate large and complex tissues at high resolution.
Check out our latest @biorxivpreprint https://doi.org/10.1101/2023.03.28.534574
A short summary thread about the software tool:🧵(1/9)
The protocol provides the step-by-step experimental methods for our @eLife paper "3D cell neighbour dynamics in growing pseudostratified epithelia" https://elifesciences.org/articles/68135.
See also our Tweetorial:
---
RT @DagmarIber
How many neighbours does an epithelial cell have? And how stable are these interactions over time? #epithelialcell #topology #LSFM #SPIM
Check out our latest pre-print: https://www.biorxiv.org/content/10.…
https://twitter.com/DagmarIber/status/1368806675495452672

3D cell neighbour dynamics in growing pseudostratified epithelia
The 3D shape and dynamic organization of epithelial cells are far more complex than previously believed, but can be explained with simple physical principles based on lateral surface energy minimisation.
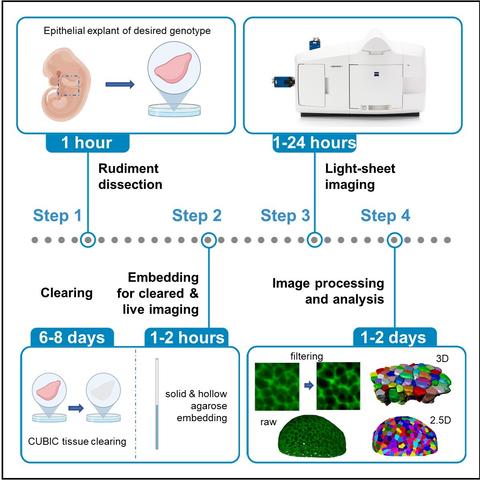
As theoretician, I am excited to share the 1st experimental protocols paper from my lab
"Time-lapse and cleared imaging of mouse embryonic lung explants to study 3D cell morphology and topology dynamics"
https://star-protocols.cell.com/protocols/2552
Thx @starprotocols for inviting this protocol!!
Cell Press: STAR Protocols
Web site created using create-react-app
Pleased to see Franziska Lampart's & Nick Doumpas' review now out:
Organoids in high-throughput and high-content screenings
https://doi.org/10.3389/fceng.2023.1120348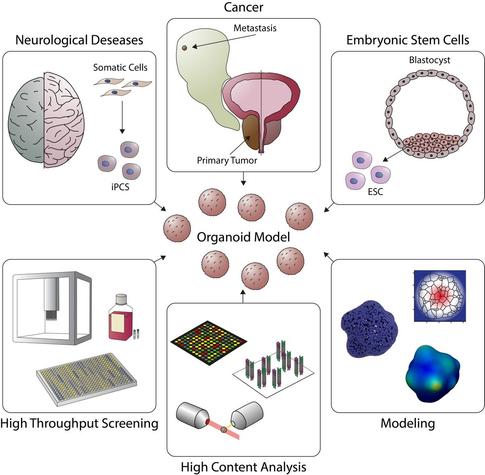

Organoids in high-throughput and high-content screenings
Organoids are self-organized three-dimensional (3D) multicellular tissue cultures which derive from cancerous and healthy stem cells, sharing a highly similarity to the corresponding in vivo organs. Since their introduction in 2009, they have emerged as a valuable model for studying early embryogenesis, organ and tissue development, as well as tools in drug screening, disease modeling and personalized therapy. Organoids can now be established for various tissues, including brain, retina, thyroid, gastrointestinal, lung, liver, pancreas, and kidney. These micro-tissues resemble the native organ in terms of gene expression, protein expression, tissue architecture and cell-cell interactions. Despite the success of organoid-based research and the advances in patient-derived organoid culture, important challenges remain. In this review, we briefly showcase the evolution from the primary 3D systems to complex, multilayered 3D structures such as assembloids, gastruloids and ETiX embryoids. We discuss current developments in organoid research and highlight developments in organoid culturing systems and analysis tools which make organoids accessible for high-throughput and high-content screening. Finally, we summarize the potential of machine learning and computational modeling in conjunction with organoid systems.