As theoretician, I am excited to share the 1st experimental protocols paper from my lab
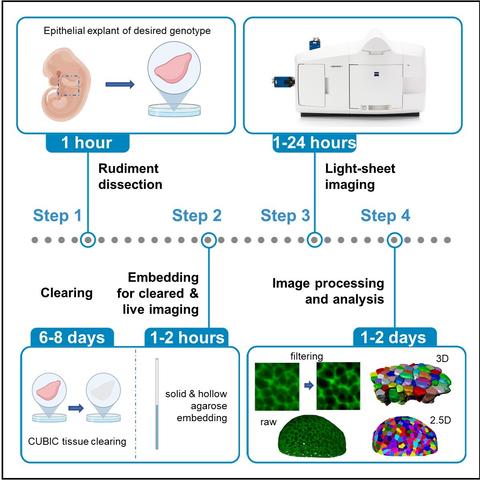
"Time-lapse and cleared imaging of mouse embryonic lung explants to study 3D cell morphology and topology dynamics"
https://star-protocols.cell.com/protocols/2552
Thx @starprotocols for inviting this protocol!!