Thrilled to share our
#OpenAccess online resource "An Introduction to Medicinal Chemistry & Molecular Recognition" on @OpenLibraryON! We designed this resource to provide budding medicinal chemists with the knowledge & tools to navigate small molecule drug discovery!
#OERhttps://ecampusontario.pressbooks.pub/medicinalchemistry/An Introduction to Medicinal Chemistry & Molecular Recognition – Simple Book Publishing
Check out our preprint on an
#AI-powered
#ligandability prediction platform for bottom-up analysis of
#chemoproteomic data to identify
#covalent binding sites, including cryptic sites, for the discovery of novel drug targets.
#proteomics #MachineLearning https://chemrxiv.org/engage/chemrxiv/article-details/658de2e266c13817293b8174

Chemical Proteomics-based Target Prioritization through a Residue Agnostic Ligandability Assessment Platform
The landscape of drug discovery is undergoing a transformative phase with the influx of structural biology and omics data. Identifying optimal drug targets amid this data surge presents a multifaceted challenge. Covalent inhibitors, once undervalued, now hold substantial promise, especially targeted covalent inhibitors (TCIs), effectively engaging 'undruggable' proteins and overcoming resistance mechanisms. Existing ML software can proficiently model covalent ligands but lack comprehensive utility across large chemoproteomics sites. Challenges persist in predicting and assessing cryptic ligandable sites and sites beyond cysteine, demanding advanced computational tools. As cysteine-ligandable proteins represent only ~20% of the quantifiable proteome, there is a requirement for ligandability mapping of other nucleophilic amino acids. This study introduces a pioneering computational pipeline leveraging an AI-based ligandable predictor for meticulous evaluation of chemical proteomics-based reactive sites. The pipeline offers a scalable framework to assess covalent ligandability on a large scale, filter out improbable hits and systematically evaluate potential drug targets. Our work addresses covalent drug design challenges through a pipeline that fills crucial gaps in predicting cryptic ligandable and covalent sites in addition to cysteines to foster more efficient drug discovery methodologies.
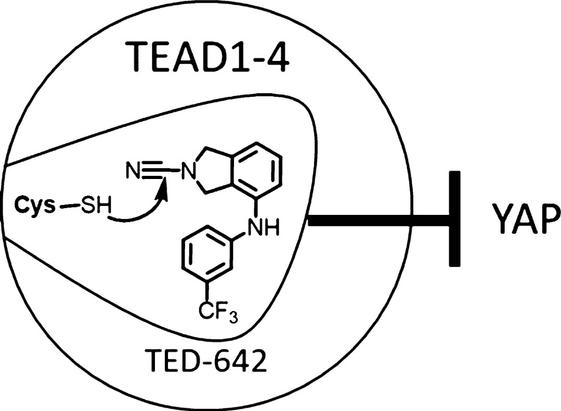
Interesting work by Samy Meroueh @IUMedSchool on the development and characterization of reversible
#covalent cyanamide antagonists of TEAD-YAP1 that engage a conserved cysteine palmitoylation site in
#TEAD.
https://pubs.acs.org/doi/abs/10.1021/acs.jmedchem.2c01189We're a
#medicinalchemistry lab in the Centre for Medicinal Chemistry at the University of Toronto Mississauga. Our lab is a dynamic research environment with pharma-quality facilities. Research translation is a central goal of our work, and several companies have sprung out of technologies developed in our lab!
https://utm.utoronto.ca/cmc/Centre for Medicinal Chemistry | Centre for Medicinal Chemistry