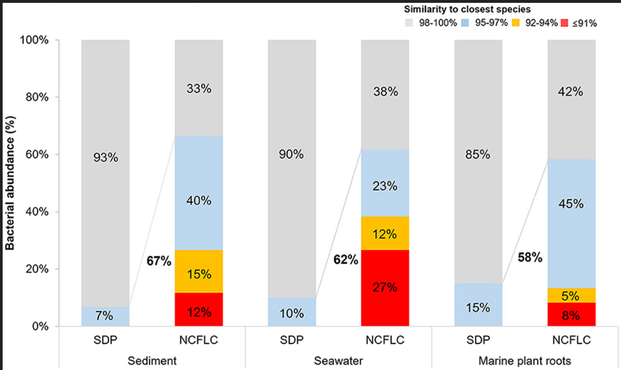
Here they do a dilution to extinction experiment where they purposefully screen for wells that do *not* grow on solid media. What do they find? A huge enrichment in undercultivated microbes: Acidobacteria, Verrucomicrobia, Armatimondaetes!
Really simple, but seems really effective!
https://www.frontiersin.org/articles/10.3389/fmicb.2023.1194466/full
A targeted liquid cultivation method for previously uncultured non-colony forming microbes
A large number of microbes are not able to form colonies using agar-plating methods, which is one of the reasons that cultivation based on solid media leaves the majority of microbial diversity in the environment inaccessible. We developed a new Non-Colony-Forming Liquid Cultivation method (NCFLC) that can selectively isolate non-colony-forming microbes that exclusively grow in liquid culture. The NCFLC method involves physically separating cells using dilution-to-extinction (DTE) cultivation and then selecting those that could not grow on a solid medium. The NCFLC was applied to marine samples from a coastal intertidal zone and soil samples from a forest area, and the results were compared with those from the standard direct plating method (SDP). The NCFLC yielded fastidious bacteria from marine samples such as Acidobacteriota, Epsilonproteobacteria, Oligoflexia, and Verrucomicrobiota. Furthermore, 62% of the isolated strains were potential new species, whereas only 10% were novel species from SDP. From soil samples, isolates belonging to Acidobacteriota and Armatimonadota (which are known as rare species among identified isolates) were exclusively isolated by NCFLC. Colony formation capabilities of isolates cultivated by NCFLC were tested using solid agar plates, among which approximately one-third of the isolates were non-colony-forming, approximately half-formed micro-colonies, and only a minority could form ordinary size colonies. This indicates that the majority of t...