SMT (@[email protected]...

SMT (@[email protected])
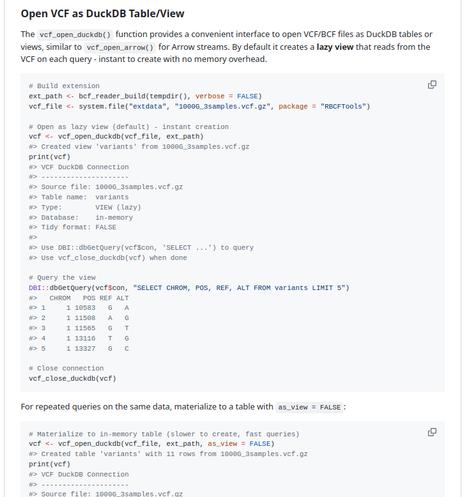
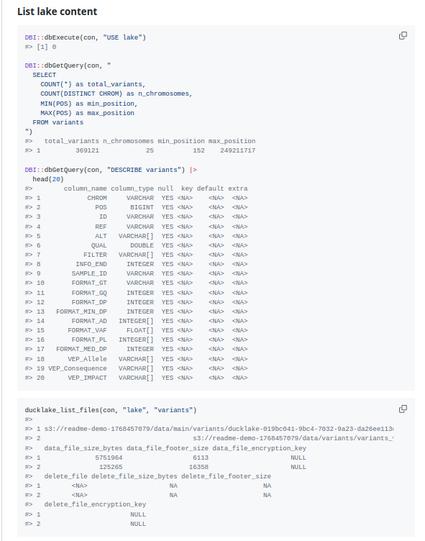
Attached: 4 images #Genomics #Bioinformatics Release of duckhts: #htslib based #Duckdb Extension for High Throughput Sequencing File Formats https://duckdb.org/community_extensions/extensions/duckhts Allied Self contained #Rstats package Rduckhts : https://rgenomicsetl.r-universe.dev/Rduckhts For now the lifecycle of the APIs (Duckdb C API extension and R package) are "experimental". They are mostly targeted to some ETL use cases, but will add some some useful scalar and aggregation functions including Heng Li's C Genomic ranges Feedback and testing welcome !