And another ding for #VScode/#positron as a package developer: I cannot run #roxygen examples (at least I can still do that in #Nvim (or a simulacrum of it as long as I'm careful: https://github.com/R-nvim/R.nvim/issues/106))
[edit: I have to run the roxygen comments line-by-line: https://github.com/posit-dev/positron/issues/1410#issuecomment-1787438498]
https://github.com/posit-dev/positron/issues/2077
https://github.com/REditorSupport/vscode-R/issues/147
Error when sending code to the console when there are roxygen2-type comments · Issue #106 · R-nvim/R.nvim
If there are roxygen2-type comments (preceded by #') and these are part of a selection being sent to the R console, there will be an error. E.g., if you have MyFunction = function(x) { print(x) } #...
 I was thinking to switch over to #VScode or #positron, but having to learn new shortcuts for everything make me sad.
I was thinking to switch over to #VScode or #positron, but having to learn new shortcuts for everything make me sad.The {document} #rstats 📦 creates roxygen2-style documentation for functions that aren't (yet) part of an R package.
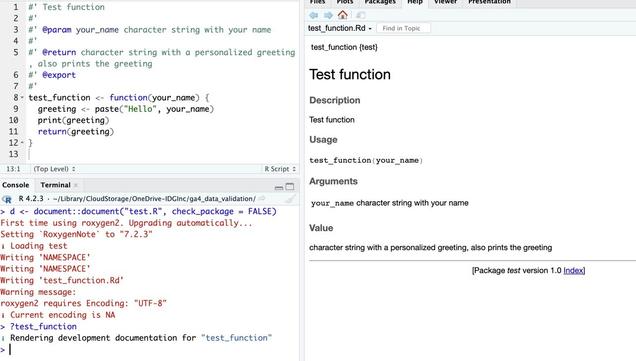
In the screenshot below, top left shows a test.R file with a function called test_function , including roxygen definitions. After running
d <- document("test.R")
I can use
?test_function
to read the help file as if it were part of a package.
By Andreas Dominik Cullmann
https://cran.r-project.org/web/packages/document/vignettes/Introduction_to_document.html
Recently ran into a disturbing #ChatGPT result: I asked it to translate a #python function written by Mathieu Daëron (https://github.com/mdaeron/D47crunch) to #R, using the #tidyverse and a #tibble. I had done this manually for my #rpackage https://github.com/isoverse/clumpedr/ and wanted to see how they would compare.
It returned my code literally, including TODO notes, comments, and #roxygen skeleton. But it did not say where it got the code from, even when pressed. My R package is on #github under #GPL 3. Thoughts?
GitHub - mdaeron/D47crunch: Python library for processing and standardizing carbonate clumped-isotope analyses, from low-level data out of a dual-inlet mass spectrometer to final, “absolute” Δ47 and Δ48 values with fully propagated analytical error estimates.
Python library for processing and standardizing carbonate clumped-isotope analyses, from low-level data out of a dual-inlet mass spectrometer to final, “absolute” Δ47 and Δ48 values with fully prop...