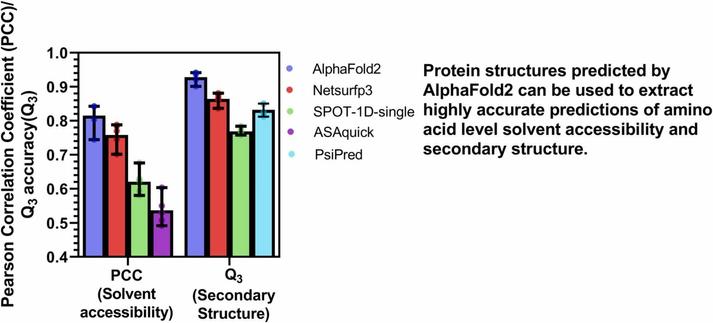
🧩 What happens when we put AlphaFold2’s predictions to the ultimate test—across species and structure types?
🔗 Comprehensive assessment of AlphaFold’s predictions of secondary structure and solvent accessibility at the amino acid-level in eukaryotic, bacterial and archaeal proteins. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2025.05.047
📚 CSBJ: https://www.csbj.org/
#AlphaFold #Bioinformatics #StructuralBiology #AlphaFold2 #ProteinStructure #ProteinPrediction #AI
Nice study from @teresa_omeara & @maom in @biorxivpreprint
They used protein prediction to explore the poorly annotated Candida auris genome, and then used protein design to replace endogenous proteins with de novo designed ones.
https://www.biorxiv.org/content/10.1101/2025.05.14.654010v1
#StructuralBiology #AlphaFold2 #proteinPrediction

Deep homology and design of proteasome chaperone proteins in Candida auris
A central tenet of biology is that protein structure mediates the sequence-function relationship. Recently, there has been excitement about the promise of advances in protein structure modeling to generate hypotheses about sequence-structure-function relationships based on successes with controlled benchmarks. Here, we leverage structural similarity to identify rapidly evolving proteasome assembly chaperones and characterize their function in the emerging fungal pathogen Candida auris. Despite the large sequence divergence, we demonstrate conservation of structure and function across hundreds of millions of years of evolution, representing a case of rapid neutral evolution. Using the functional constraints on structure from these naturally evolved sequences, we prospectively designed de novo chaperones and demonstrate that these artificial proteins can rescue complex biological processes in the context of the whole cell.
### Competing Interest Statement
The authors have declared no competing interest.
National Institutes of Health, https://ror.org/01cwqze88, R35GM147894, U19AI181767, T32GM149391, R35GM151129, 5F32CA261115, R35 GM118073
University of MichiganAnn Arbor, https://ror.org/00jmfr291, Postdoctoral pioneer fellowship
🔬 Exciting showdown alert! 🌟 Dive into the world of protein structure prediction with
#RosettaFold All Atom vs.
#AlphaFold - two powerhouses revolutionizing the field. 🧬
https://www.eliza-ng.me/post/sidentifiedfrom_4/#ProteinPrediction #Bioinformatics #BlogPost
The Protein Prediction Showdown: RosettaFold All Atom vs. AlphaFold - Breaking Boundaries in Protein Structure Prediction
A new protein structure prediction model released by David Baker’s laboratory is making waves in the scientific community, challenging the dominance of DeepMind’s AlphaFold. The model, named RosettaFold All Atom, not only predicts protein structures but also incorporates bound DNA and ligands. What sets this model apart is its open-source nature, allowing for greater accessibility and collaboration within the scientific community.
The rivalry between DeepMind’s AlphaFold and Baker’s laboratory model adds an exciting dynamic to the field of protein structure prediction.
@cmdcolin
I use Alphafold (actually colabfold) and am happy to answer questions, but I'm just a user. There are people with more expertise than me here.
Hashtags might find the right people
#StructurePrediction #alphafold #alphafold2 #colabfold #ProteinStructure #StructuralBiology #ProteinPrediction #protein #bioinformatics
Note in point.
What does it mean when only your rank 1 model suggests an interaction between two proteins with alphafold multimer? Is this garbage?
#AlphaFoldMultimer
#AlphaFold
#AlphaFold2
#ProteinPrediction
#StructurePredictions
#ProteinPredictionHelp