… ligand–protein BSA of all molecular glue ternary complex structures. The asymmetry of interactions between the molecular glue and its protein binding partners can guide the understanding of their ternary structure formation.
These analysis should help the field when thinking about how to identify degrading and non-degrading molecular glues in a prospective way. Thank you to co-authors Huan Rui, Jaeki Min, Kate Ashton, and Connie Wang.
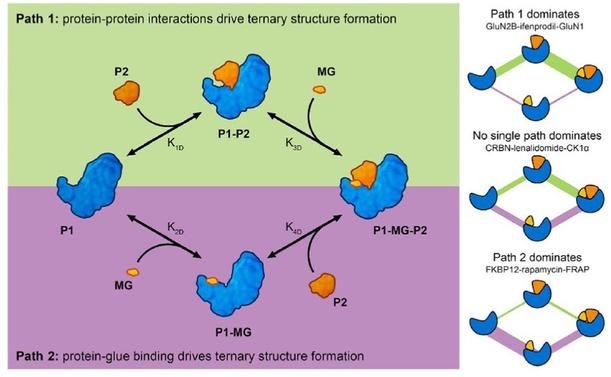
#TPD #GLUEs #InducedProximityAnalysis of molecular glue-induced ternary structures indicate that there are 2 main pathways of how ternary structure is formed; Path 1) a protein–protein binary interface is formed first and then the complex incorporates a small molecule ligand at the interface or Path 2) small molecule binds to a protein partner first, which leads to altered protein surface properties, much like the effects of post-translational modification, making this surface available for association with another protein
We then analyze the molecular features of these protein-protein interactions, including buried surface area (BSA), interface amino acid preference profiles, residue pair preference profiles, and …
We summarize >100 molecular glue-induced ternary complexes, Group 1) features interactions between proteins with well-folded domains, Group 2) one of the binding partners is a stretch of residues that contains a specific pattern for binding based on interaction mode.
New Today! Induced Proximity group at @Amgen publish in @rsc_chembio on Protein–protein interfaces in molecular glue-induced ternary complexes: classification, characterization, and prediction.
https://tinyurl.com/5n95hz79. Main conclusions… 1/thread
… need to wrangle molecule geometries, etc, to achieve productive ubiquitination by unnatural ligase for TPD therapeutics. And thus less chemistry = quicker development. Great example of ever expanding induced proximity modalities.
#TPD #proximity #PROTAC #ubiquitin /end
… substrate dependent. So this remains a big open question of how generalizable the concept is. 5) No proteomics shown as to potential impact on binding PSMD2. Presumably the macrocycle is substiocheometric but future studies with small molecules will need to be careful. 6) No…
… evidence provided as to whether BRD4 still needed to be ubiquitinated. This remains a question of mine as it could impact maximize kinetics if reliant on some level of endogenous Ub before degradation. 7) Implications here are large. Skipping ligase would likely mean less…
… 26S proteasome to recruit targets. PSMD2 is natural receptor of ubiquitinated proteins and close to ATPase pore for unraveling substrates. 2) Very impressive use of mRNA display to identify macrocyclic peptides that bind PSMD2. 3) As expected, macrocycles are not very…