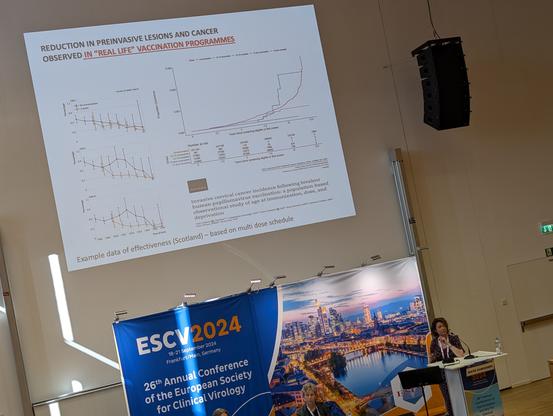
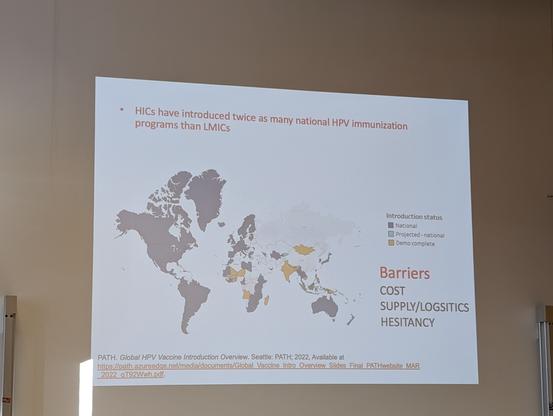
Every time I hear about HPV vaccines & cervical cancer I want to shout from the rooftops - it's incredible to live in a time where we can prevent this devastating disease!
But as Kate Cuschieri highlights, we have to tackle this globally & reduce inequity in access! #ESCV2024


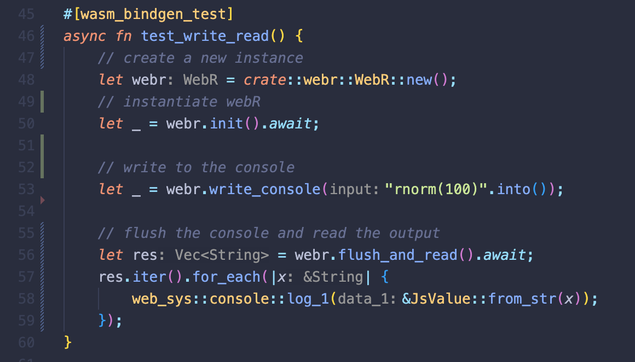
 package authors.
package authors.