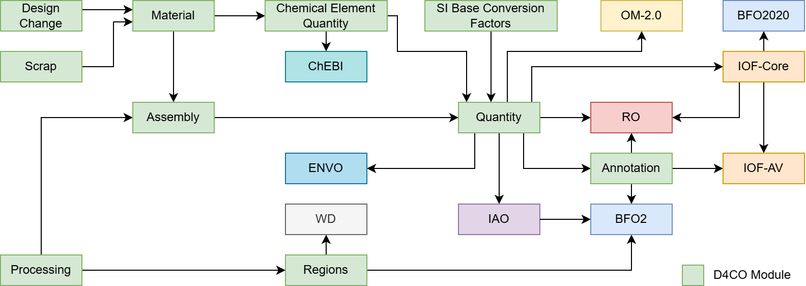
Already yesterday, I presented at the ESWC 2026 Pre-Conference in the Knowledge Graphs for Sustainability Workshop (KG4S) the Design-for-Circularity Ontology (D4CO). Related artifacts are available online.
🔗 Ontology: https://w3id.org/d4co
🧑💻 Repository: https://gitlab.com/dlr-dw/d4co
📄 Paper Pre-Print: https://elib.dlr.de/224130/