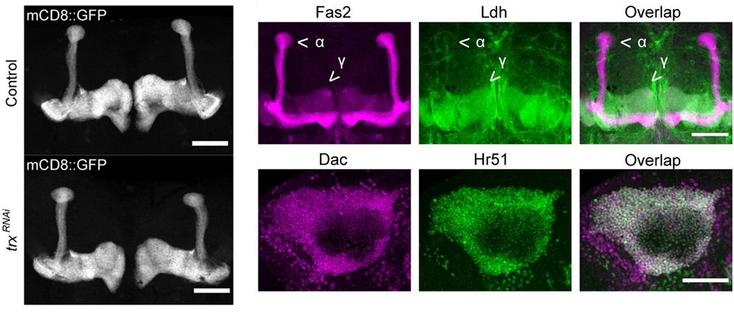
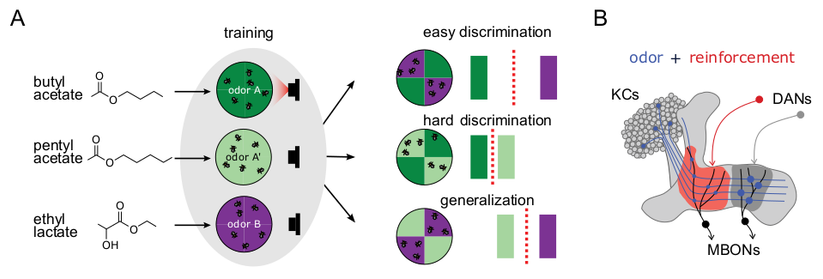
Synapses in the fly brain learning and memory centre, the mushroom body:
"Quantitative characterization of the pattern of Brp clusters across multiple individuals revealed cell-type-dependent synapse heterogeneity and stereotypy. Furthermore, we discovered previously unidentified sub-compartmental synapse configuration and its regulation by cAMP signaling."
From:
"High-throughput synapse profiling reveals cell-type-specific spatial configurations in the fly brain", by Wu et al. (Tanimoto lab) 2025
https://www.biorxiv.org/content/10.1101/2024.12.02.626511v4
High-throughput synapse profiling reveals cell-type-specific spatial configurations in the fly brain
Characterization of intracellular synapse heterogeneity aides to understand the intricate computational logic of neuronal circuits. Despite recent advances in connectomics, the spatial patterns of synapses and their inter-individual variability remain largely unknown. Using directed split-GFP reconstitution, we achieved visualization of endogenous Bruchpilot (Brp) proteins, the presynaptic active zone (AZ) scaffold, in a cell-type-specific manner. By developing a high-throughput quantification pipeline, we profiled AZ structures in identified neurons of the mushroom body circuit, where intracellular synaptic pattern is crucial due to its compartmentalized connectivity. Quantitative characterization of the pattern of Brp clusters across multiple individuals revealed cell-type-dependent synapse heterogeneity and stereotypy. Furthermore, we discovered previously unidentified sub-compartmental synapse configuration and its regulation by cAMP signaling. These results of synapse profiling uncovered multi-layered organizations of AZs, ranging from neighboring synapses to consistent patterns across individuals. Teaser High-throughput profiling of single active zones revealed previously unidentified multi-layered synaptic organizations. ### Competing Interest Statement The authors have declared no competing interest.