【💡Editor's Choice】
Volatilization-related ammonia and fossil fuel nitrogen oxides emission sources are the major contributors to #AtmosphericNitrogenDeposition in the Mengshan area according to the #Mosses record.
#EnvironmentalChanges | #StableNitrogenIsotope | #Biomonitoring
https://doi.org/10.1093/jpe/rtaf039
Since 1994, eight bivalve mussels have been monitoring water quality in Poznań, Poland.
https://www.zmescience.com/ecology/poznan-mussel-water-plants-892524/ Electrical contacts are attached to them with hot glue. If the water is clean, their shells remain open, but when four mollusks close simultaneously, the system automatically shuts off the water supply to the entire city.
#WaterQuality #BioMonitoring #Mussels
In Poznan, Poland, eight clams get to decide if people in the city get water or not
They might be small, but these mussels shoulder tremendous responsibility.
✳️ #Mosses record historical variations of #AtmosphericNitrogenDeposition in a mountain area of northern China
Results:
1️⃣ 2012~2018: moss nitrogen(N) contents ⬆️, moss N isotope values ⬇️;
2️⃣ 2012~2022: total N deposition fluxes ⬆️.
Main sources:
1️⃣ volatilization-related ammonia
2️⃣ fossil fuel N oxides emission
#EnvironmentalChanges | #StableNitrogenIsotope | #Biomonitoring
https://doi.org/10.1093/jpe/rtaf039
Biomonitoring Humain Wallon (BMH-Wal) pour estimer l'exposition interne et globale des personnes aux substances (polluants, substances chimiques, pesticides, métaux...).
https://environnement.sante.wallonie.be/biomonitoring-wallon
#biomonitoring #wallon #pfas
Biomonitoring Humain Wallon (BMH-Wal) - Portail Environnement-Santé
Portail environnement santé de la Wallonie

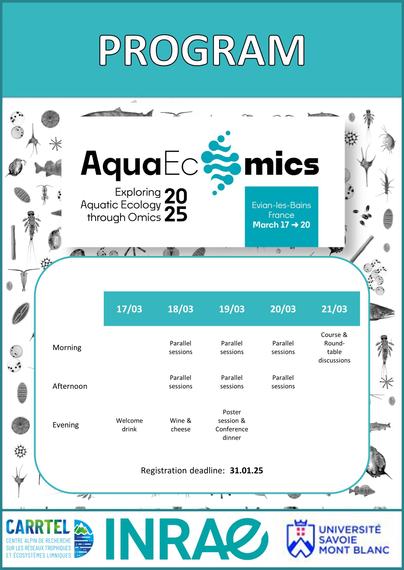
Program - AquaEcOmics
Here is a preliminary overview of the AquaEcOmics program :
Please note that th...
**Proficiency testing and cross-laboratory method comparison to support standardisation of diatom DNA metabarcoding for freshwater biomonitoring**
https://mbmg.pensoft.net/article/133264
#DNAmetabarcoding #standardization #freshwater #biomonitoring

Proficiency testing and cross-laboratory method comparison to support standardisation of diatom DNA metabarcoding for freshwater biomonitoring
DNA metabarcoding of benthic diatoms has been successfully applied for biomonitoring at the national scale and can now be considered technically ready for routine application. However, protocols and methods still vary between and within countries, limiting their transferability and the comparability of results. In order to overcome this, routine use of DNA metabarcoding for diatom biomonitoring requires knowledge of the sources of variability introduced by the different steps of the procedure. Here, we examine how elements of routine procedures contribute to variability between European laboratories. A set of four experiments were performed focusing on DNA extraction and PCR amplification steps to evaluate their reproducibility between different laboratories and the variability introduced by different protocols currently applied by the scientific community. Under the guidance of a reference laboratory, 17 participants from 14 countries performed DNA extraction and PCR amplification in parallel, using the same fixed protocol and their own choice of protocol. Experiments were performed by each participant on a set of standardised DNA and biofilm samples (river, lake and mock community) to investigate potential systematic and random errors. Our results revealed the successful transferability of a protocol amongst labs and a highly similar and consistent ecological assessment outcome obtained regardless of the protocols used by each participant. We propose an “all for one but prove them all” strategy, suggesting that distinct protocols can be used within the scientific community, as long as their consistency is be proven by following minimum standard requirements.
"Proficiency testing and cross-laboratory method comparison to support standardisation of diatom DNA metabarcoding for freshwater biomonitoring"
https://mbmg.pensoft.net/article/133264
#DNAmetabarcoding #standardization #ProficiencyTesting #biomonitoring

Proficiency testing and cross-laboratory method comparison to support standardisation of diatom DNA metabarcoding for freshwater biomonitoring
DNA metabarcoding of benthic diatoms has been successfully applied for biomonitoring at the national scale and can now be considered technically ready for routine application. However, protocols and methods still vary between and within countries, limiting their transferability and the comparability of results. In order to overcome this, routine use of DNA metabarcoding for diatom biomonitoring requires knowledge of the sources of variability introduced by the different steps of the procedure. Here, we examine how elements of routine procedures contribute to variability between European laboratories. A set of four experiments were performed focusing on DNA extraction and PCR amplification steps to evaluate their reproducibility between different laboratories and the variability introduced by different protocols currently applied by the scientific community. Under the guidance of a reference laboratory, 17 participants from 14 countries performed DNA extraction and PCR amplification in parallel, using the same fixed protocol and their own choice of protocol. Experiments were performed by each participant on a set of standardised DNA and biofilm samples (river, lake and mock community) to investigate potential systematic and random errors. Our results revealed the successful transferability of a protocol amongst labs and a highly similar and consistent ecological assessment outcome obtained regardless of the protocols used by each participant. We propose an “all for one but prove them all” strategy, suggesting that distinct protocols can be used within the scientific community, as long as their consistency is be proven by following minimum standard requirements.
Excited to announce the preliminary program for the upcoming AquaEcOmics symposium. Registration closes on the 31th of January so it is not yet too late to secure your attendance.
https://aquaecomics.symposium.inrae.fr/#aquaecomics2025 #eDNA #omics #metagenomics #metabarcoding #dPCR #marine #freshwater #ecology #biomonitoring