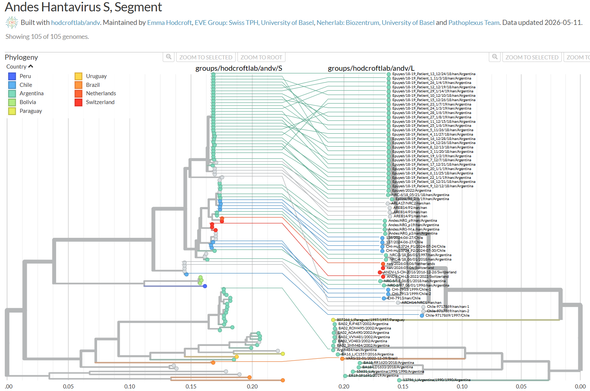
Nextstrain #ANDV trees updated with a few improvements:
- 2 outbreaks marked
- host scientific/common name added
- province now visible on map
- date colouring added
Here's a quick tour of how to change colours, geography & names!
Links to trees here (scroll down):
https://nextstrain.org/groups/hodcroftlab