Come see our deeper dive into human chromatin in situ at
#cellbio2023.
#teamtomo study led by Jon Chen and Tingsheng Liu. Poster P2005, Board B281, Monday Dec 4. Preprint:
https://www.biorxiv.org/content/10.1101/2023.07.31.551204v1.fullThe nanoscale similarities and differences between G1 and metaphase chromatin are summarized in tables. Raw cryoET VPP data will be shared at EMPIAR. Now we need to figure out how cells control nucleosome structure and packing 🤔
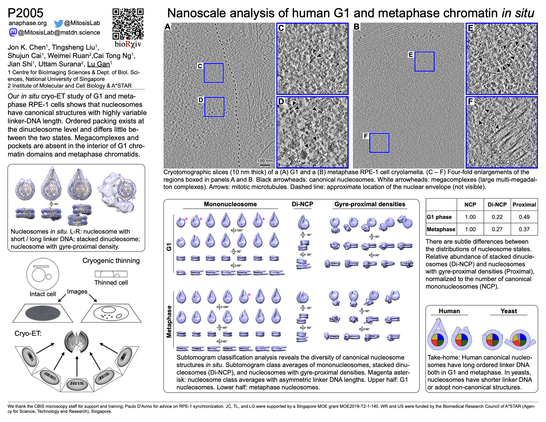
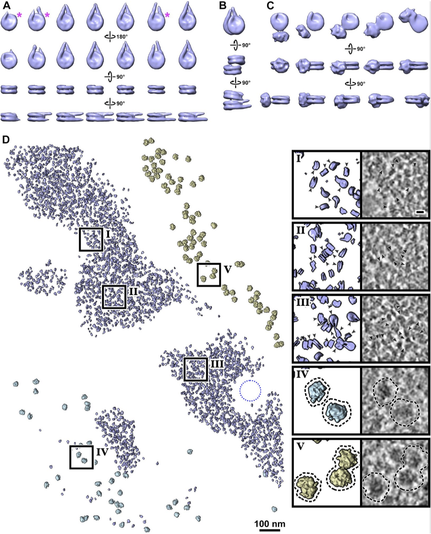
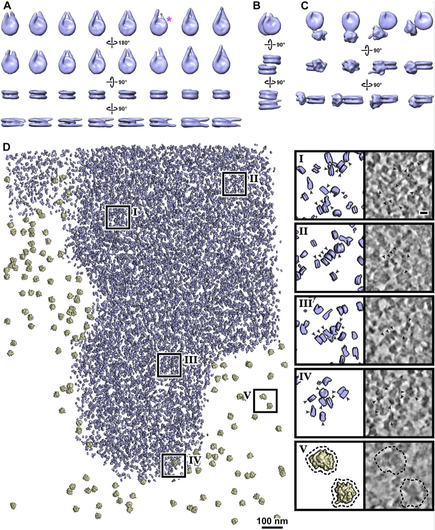
We find diverse canonical nucleosome structures in both G1 and metaphase. These include mononucleosomes, ordered stacked dinucleosomes, and nucleosomes with gyre-proximal (albeit heterogeneous) densities. Nucleosomes have variable linker-DNA length and structure.
Happy to share our in situ cryoET analysis of human chromosomes, in both G1 and metaphase. It's the work of Jon Chen , Tingsheng Liu, Shujun Cai, Cai Tong Ng, Jian Shi, and our collaborators Weimei Ruan and Uttam Surana.
https://www.biorxiv.org/content/10.1101/2023.07.31.551204v1Our non-canonical nucleosome study has been updated. tl;dr - lots of VPP
#cryoET; canonical nucleosomes account for < 10% of the total expected in situ. Work of Zhi Yang Tan, Shujun Cai, Alex Noble, and Jon Chen.
https://www.biorxiv.org/content/10.1101/2021.04.04.438362v2Happy to share Claris Chong's review on non-canonical nucleosome structures, now published at Chromosoma!
https://link.springer.com/article/10.1007/s00412-023-00791-w

Are extraordinary nucleosome structures more ordinary than we thought? - Chromosoma
The nucleosome is a DNA–protein assembly that is the basic unit of chromatin. A nucleosome can adopt various structures. In the canonical nucleosome structure, 145–147 bp of DNA is wrapped around a histone heterooctamer. The strong histone-DNA interactions cause the DNA to be inaccessible for nuclear processes such as transcription. Therefore, the canonical nucleosome structure has to be altered into different, non-canonical structures to increase DNA accessibility. While it is recognised that non-canonical structures do exist, these structures are not well understood. In this review, we discuss both the evidence for various non-canonical nucleosome structures in the nucleus and the factors that are believed to induce these structures. The wide range of non-canonical structures is likely to regulate the amount of accessible DNA, and thus have important nuclear functions.
It's great to see that a popular
#cryoEM software package is now a benchmark on a major enterprise hardware review website:
https://www.phoronix.com/review/intel-xeon-platinum-8490h/6
Intel Xeon Platinum 8490H "Sapphire Rapids" Performance Benchmarks Review
We're looking forward to presenting the work of @
[email protected], @
[email protected], and @
[email protected], which addresses the impact (or not) of transcription on nucleosome structure in situ. Please see our
#CellBio2022 Dec 5 microsymposium talk in Rm 207B and Dec 6 poster B283!