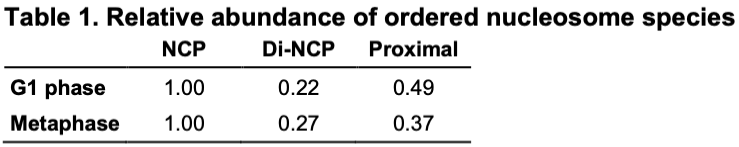
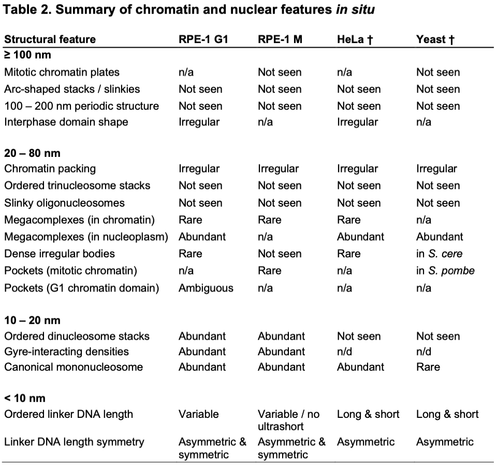
Happy to share our in situ cryoET analysis of human chromosomes, in both G1 and metaphase. It's the work of Jon Chen , Tingsheng Liu, Shujun Cai, Cai Tong Ng, Jian Shi, and our collaborators Weimei Ruan and Uttam Surana.
https://www.biorxiv.org/content/10.1101/2023.07.31.551204v1
https://www.biorxiv.org/content/10.1101/2023.07.31.551204v1