https://journals.biologists.com/jcs/article/136/24/jcs261594/337755/Positive-directions-from-negative-results
This Essay is associated with a wild cherry tree planted in the Forest of Biologists @Co_Biologists https://forest.biologists.com/landscape/?id=86128
Positive directions from negative results
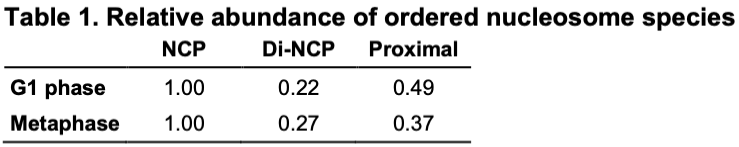
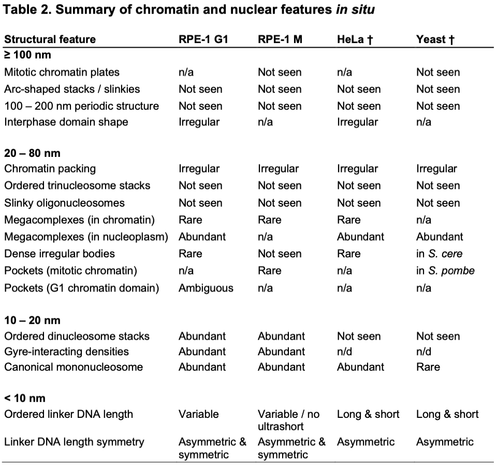
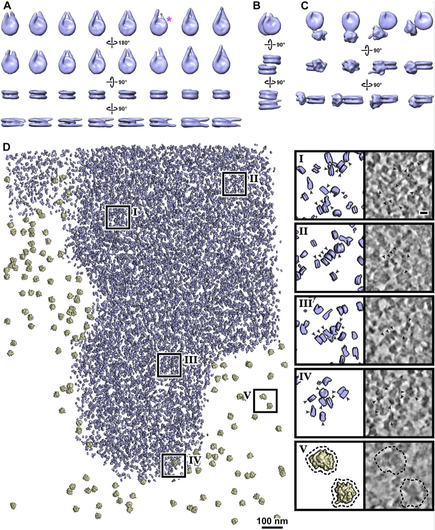
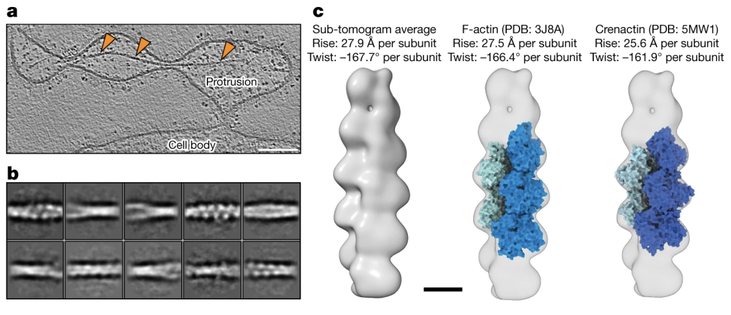
Studies often produce ‘negative’ results: they are well-controlled and reproducible, but they do not produce the expected results. And yet positive results represent the vast majority of published literature (Fanelli, 2012), meaning that negative results are often unreported. Nevertheless, negative results are important because they expose a fundamental knowledge gap, and the eventual resolution to this knowledge gap might have an immensely positive impact on science and the world in general. In this essay, I will use three brief stories – one that gained fame due to the recent COVID-19 pandemic and two from my field of chromatin biology – to convince you that negative results drive exciting new research directions, and that just because a result contradicts expectation, it does not mean that it is a negative result.It is estimated that more than two billion doses of the new RNA-based COVID-19 vaccines have been administered (https://ourworldindata.org/grapher/covid-vaccine-doses-by-manufacturer). This technology is significant because it has many advantages over traditional DNA and viral-vector-based vaccines (Pardi et al., 2018). Many labs have experimented with RNA as a gene therapy tool but were unsuccessful in adapting it to the clinic for nearly 30 years (Hou et al., 2021; Sahin et al., 2014). In other words, RNA therapeutics were beset by many negative results for 30 years! One of the key barriers to the use of RNA as a vaccine is that it triggers the innate immune response (Karikó et al., 2004; Weissman et al., 2000), meaning that dangerous side effects can result. A key breakthrough, which was recognized by the 2023 Nobel Prize in Medicine, was the discovery that unmodified RNA was the structure detected by the innate immune system (Karikó et al., 2005). This ‘eureka’ moment suggested that RNA therapeutics – including the COVID-19 vaccines that were invented 15 years later – could be made safer by the incorporation of modified RNA. In other words, the years of research into RNA's interaction with the innate immune response, driven by the motivation to resolve these negative results, eventually led to the seminal ‘positive’ result that suggested how RNA may be safely used as a clinical reagent. This RNA therapeutics story illustrates how the identification of a knowledge gap (low tolerance of unmodified RNA) and its ultimate resolution helped make possible a world-changing biomedical advance.Chromatin is built from chains of basic units called nucleosomes, 10-nm-wide, 6-nm-thick cylindrical complexes within which DNA is wrapped. Early in vitro studies of isolated chromatin led to a model of higher-order chromatin structure where nucleosomes pack as crystalline 30-nm-wide fibers (Finch and Klug, 1976). Around a decade later, pioneers of chromatin structural cell biology studied chromatin by analyzing cryo-electron microscopy (cryo-EM) images of cryosections from mammalian cells (McDowall et al., 1986). They found a negative result – the images were consistent with a model in which in situ chromatin is liquid-like (irregular) and not crystalline. This finding remained under-appreciated until two decades later, when the (irregular) in situ structure of chromatin was revisited using a human cell line (HeLa) and with improved image analysis (Eltsov et al., 2008). This later study supported the pioneering one – there was no evidence for densely packed ordered 30-nm fibers. Subsequent studies performed by other groups in different cell lines and cell cycle states, and using purified chromosomes, continued to support the original ‘negative’ result from 1986, resulting in the acceptance that chromatin is irregular in situ, not crystalline (Maeshima et al., 2019). Decades of analysis using newer technologies have now converted the irregular in situ chromatin packing model from an unexpected to an expected one. Irregular chromatin now features in newer concepts of chromatin regulation, such as how chromatin can have liquid-like behavior in situ (Nozaki et al., 2023; Strickfaden et al., 2020).To achieve the resolution needed to resolve nucleosomes inside cells, structural cell biologists use a form of cryo-EM called cryo-electron tomography (cryo-ET) as the main tool. Cryo-ET generates 3D models of unique objects (e.g. cells) at ∼4 nm resolution. The high-resolution nucleosome crystal structure that was determined 26 years ago is considered the ‘canonical’ and most abundant one (Luger et al., 1997). Subsequent studies of reconstituted nucleosomes (Zhou et al., 2019), including one that used recombinant yeast histones (White et al., 2001), have revealed very similar structures. These nucleosome structures therefore reinforce the expectation that nucleosomes are universally canonical. Based on this expectation, we were surprised that in our in situ cryo-ET experiments of Saccharomyces cerevisiae (budding yeast) nucleosomes, no evidence of canonical nucleosomes could be found. The finalized version of this study showed that in a popular wild-type budding yeast strain, less than 10% of the nucleosomes have the canonical structure, meaning that the vast majority (>90%) have non-canonical structures (Tan et al., 2023), which we suggested to have partial DNA detachment and variable histone orientations. This unexpected result was in stark contrast to our earlier study of a HeLa cell, which had six times more canonical nucleosomes per unit nuclear volume (Cai et al., 2018). To convince ourselves and the community that our ‘controversial’ observations were a result of cell biology (and not, for example, incorrect experimental setup or data analysis), we did large numbers of both in situ and computational controls, which were important to ensure confidence in our findings. More importantly, our non-canonical nucleosome study guided us to new research directions about what the structures of non-canonical nucleosomes are and what causes them to be so abundant in situ, highlighting the value of following up on ‘negative’ or unexpected results.It is my hope that more negative results will be published and publicized. Furthermore, I hope that, as with positive results, review articles will be published that summarize the negative results within a field. Such concerted efforts will help identify and prioritize knowledge gaps, as well as enhance future projects by precisely defining the ‘known unknowns’ (see blog article by Andrea Mantovani). Negative results should therefore be celebrated and followed up with studies to test each of the hypotheses that could explain why the result was labeled as ‘negative’ in the first place. When the most exciting of these negative results are resolved – by bridging the knowledge gap – it would represent the conversion of previously negative results into positive ones that clarify our new models of biology.I thank C. Chong for critical comments.