| Homepage | https://www.bork.embl.de/ |
| GitHub | https://github.com/grp-bork/ |
| BlueSky | https://bsky.app/profile/borklab.bsky.social |
Bork Group at EMBL Heidelberg
- 366 Followers
- 97 Following
- 143 Posts
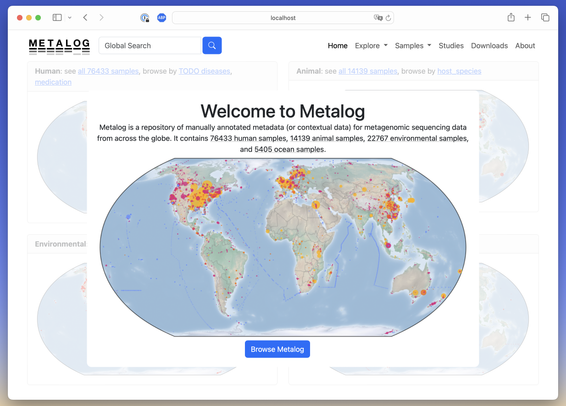
The paper by @biocs has been published in the NAR Database Issue! https://doi.org/10.1093/nar/gkaf1118
Visit the database itself on https://metalog.embl.de/ to access manually annotated contextual data for #microbiome samples around the globe. We're up to 1000 included studies as of today!
Apply until December 15th!
More info:
group homepage: https://www.bork.embl.de/
TREC: https://www.embl.org/about/info/trec/
BIOcean5D, the funding source: https://biocean5d.org/
SPIRE: https://spire.embl.de
Also, as a teaser, we plan to release out database of contextual data (a.k.a. metadata) for metagenomic samples soon! (4/4)
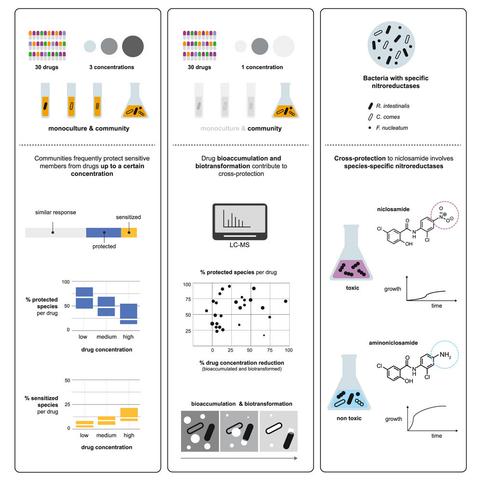
Our paper together with @TypasLab is out now! https://doi.org/10.1016/j.cell.2024.08.037 Previously available on bioRxiv, it has now been accepted in Cell. 🎉
Sarela Garcia-Santamarina and @biocs show that bacteria can be protected from & sensitized to drugs in community; that cross-protection dissipates at high drug concentrations; that biotransformation & bioaccumulation can partially explain communal protection; and that enzyme specificity & expression dictate detoxification of nitroaromatic drugs
Marisa Keller (@[email protected])
4 Posts, 54 Following, 57 Followers · Phd Candidate | Scientist working with the Microbiome 🧫 and Computational Biology 👩🏼💻 she/her | also likes music 🎶, carnival 🥳, nature 🌱 & hiking 🏔 📍Heidelberg, Germany
Probiotics and Gut Dysbiosis in Preterm Infants
The Priming Immunity at the Beginning of Life (PRIMAL) randomized clinical trial investigates if preterm infant exposure to multistrain Bifidobacteria and Lactobacillus probiotics reduces the colonization rate of multidrug-resistant organisms at day 30 of life compared with placebo.
Are you coming to #IHMC in Rome? The Bork Group is represented with three posters and two talks ("Stool #microbiome analysis and its impact on progression and decompensation of liver cirrhosis" by Peer on Monday & "The global landscape of horizontal gene transfer" by Chan Yeong Kim on Sunday). Looking forward to seeing you there!
Congratulations to Askarbek Orakov, who successfully defended his PhD at the University of Würzburg today!
Peer is dressed in a Kazakh traditional “Shapan”, which is given to honorable people – a gift from Askarbek's and his wife's families. Thank you very much for this!
Importantly, we found that microbial signatures of each disease are strongly correlated with the altered microbial load of patients, indicating that change in microbial load, rather than the disease itself, is the main driver in shaping altered microbial profile in patients.
Our prediction tool, named Microbial Load Predictor (MLP), is freely available from here. https://microbiome-tools.embl.de/mlp/