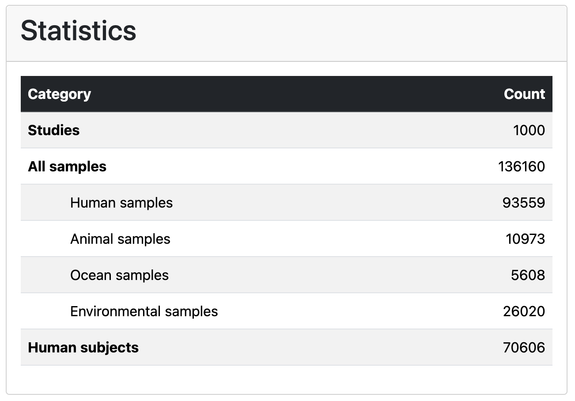
New preprint from lab: https://www.biorxiv.org/content/10.1101/2025.08.14.670324v1 describing the Metalog database of manually annotated contextual data for >110k metagenomics samples around the globe https://metalog.embl.de/
See Bluesky thread for more info: https://bsky.app/profile/biocs.bsky.social/post/3lwgfkpthjs2d