| Homepage | https://www.bork.embl.de/ |
| GitHub | https://github.com/grp-bork/ |
| BlueSky | https://bsky.app/profile/borklab.bsky.social |
Bork Group at EMBL Heidelberg
- 366 Followers
- 97 Following
- 143 Posts
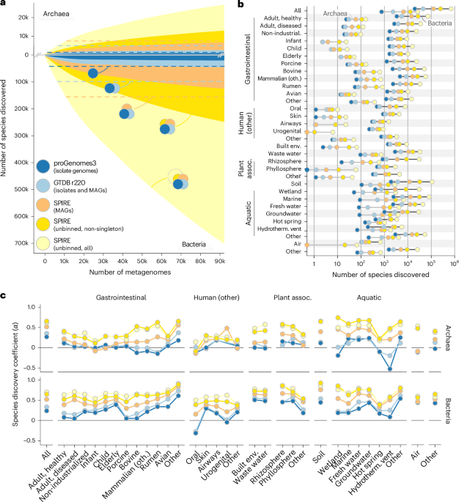
There is a lot of undescribed prokaryotic diversity in existing whole-genome metagenomic data!
https://www.nature.com/articles/s41564-026-02314-6
We predict that there are >500,000 novel bacterial and >20,000 novel archaeal species. There are at least 145 novel bacterial and 10 archaeal phylum-level clades. While according to GTDB, there are genomes for only <140,000 bacterial and <20,000 archaeal species (which corresponds to169 and 20 phyla in Bacteria and Archaea respectively).
Peer did pioneering work in so many research fields, and we aim that our research on the microbiome within the human gut and across environments will continue in his spirit. We have many ongoing projects, papers under review, and some coming out soon.
Thank you to everyone who reached out or shared what he meant to them and how he inspired them! We will keep working together, striving to fulfill Peer's scientific legacy. We hope you'll follow along!
It has been two weeks since the unexpected death of Peer Bork, and all of us in his research group are deeply missing him. His scientific vision brought us together as a team, and we are immensely grateful for the time we spent together. 🧵
https://www.embl.org/news/people-perspectives/remembering-peer-bork-obituary/
Dear #Microbiology community 🧫
2025 was a highly successful year for #nfdi4microbiota: mid-term evaluation passed with flying colors, renewal proposal submitted & defended, widely used FAIR services launched, expanded training & outreach, and strengthened collaborations across #NFDI life sciences consortia. A highlight: our Annual Conference in Cologne, bringing the community together for exchange & networking.
We wish you peaceful holidays and a great start to a healthy and successful 2026! 🎄✨
During my PhD @borklab.bsky.social, we extracted drug side effect data from package inserts and used the dataset to predict new drug targets (https://doi.org/10.1126/science.1158140) and pinpoint proteins that mediate the side effects (https://doi.org/10.1038/msb.2013.10). We also published the underlying database, but life intervened and funding changed, so SIDER http://sideeffects.embl.de/ is unfortunately gathering dust since its last update in 2015.
But there's good news: ChEMBL and OpenTargets are starting a new effort to mine the available data and produce a high-quality databases! Text-mining tools have progressed a lot in the last two decades, and I'm excited to see what can be done now.
More info https://chembl.blogspot.com/2025/12/new-position-nlp-data.html
Apply here: https://embl.wd103.myworkdayjobs.com/EMBL/job/Hinxton-Cambridgeshire/NLP-Data-Scientist-Scientific-Data-Engineer_JR2866 The position is at EBI.
Drug Target Identification Using Side-Effect Similarity
Targets for drugs have so far been predicted on the basis of molecular or cellular features, for example, by exploiting similarity in chemical structure or in activity across cell lines. We used phenotypic side-effect similarities to infer whether ...
metaTraits: https://metatraits.embl.de/ 🦠 Interactively explore 140+ 𝗺𝗶𝗰𝗿𝗼𝗯𝗶𝗮𝗹 𝗽𝗵𝗲𝗻𝗼𝘁𝘆𝗽𝗶𝗰 𝘁𝗿𝗮𝗶𝘁𝘀, harmonized & integrated from culture-derived collections 🔬 & genome-based predictions 🧬 for >2M MAGs & genomes.
See the publication at NAR: https://doi.org/10.1093/nar/gkaf1241 and the BlueSky thread by @dpodlesny https://bsky.app/profile/podlesny.bsky.social/post/3m6zj2zoptc2y
Even more resource papers from the group in the NAR Database Issue!
VIRE is a planetary-scale database of >1.7M viral genomes reconstructed from 100k+ metagenomes across diverse environments https://doi.org/10.1093/nar/gkaf1225 , led by Suguru Nishijima. Website: https://vire.embl.de/
The paper by @biocs has been published in the NAR Database Issue! https://doi.org/10.1093/nar/gkaf1118
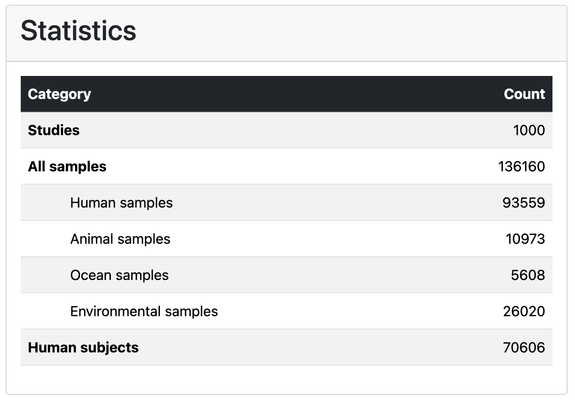
Visit the database itself on https://metalog.embl.de/ to access manually annotated contextual data for #microbiome samples around the globe. We're up to 1000 included studies as of today!