Geometry of the cumulant series in diffusion MRI
이 논문은 확산 MRI(dMRI) 신호의 누적량 텐서(cumulant tensors)의 대칭성과 불변량을 SO(3) 회전군 이론을 통해 체계적으로 분석한다. 3+18개의 회전 불변량(RICE)을 정의하여 기존 dMRI 지표들을 통합하고, 다발성 경화증 분류에서 기존 지표보다 우수한 성능을 보임을 임상 데이터로 입증했다. 또한, 1~2분 내에 주요 불변량을 측정하는 초단시간 임상 프로토콜을 설계하여 고급 dMRI의 임상 적용 가능성을 높였다. 이 연구는 dMRI 신호의 기하학적 이해를 바탕으로 머신러닝 기반 병리 및 발달 분류에 활용될 수 있다.
https://www.nature.com/articles/s41467-026-70018-w
#diffusionmri #tensoranalysis #grouprepresentation #medicalimaging #machinelearning
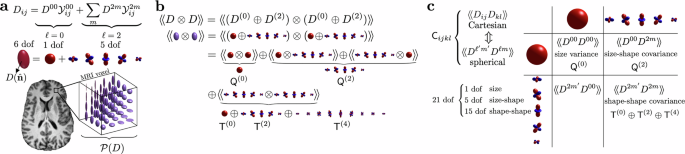
Geometry of the cumulant series in diffusion MRI - Nature Communications
Rotational symmetry defines the geometry and topology of the diffusion MRI signal, its irreducible components and their invariants. These metrics are applied to human brain MRI, improve multiple sclerosis classification, and enable faster scans.