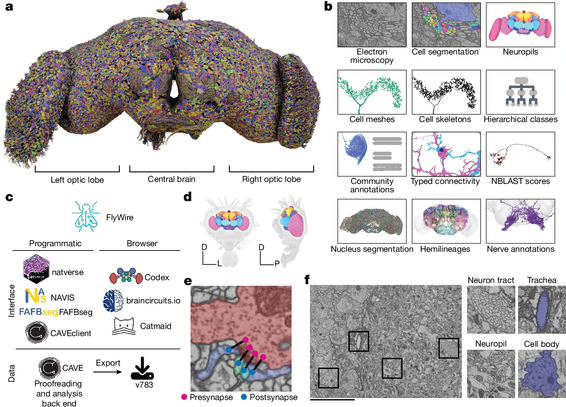
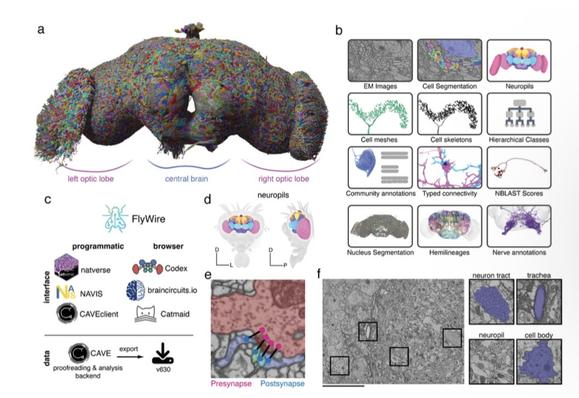
Whole adult fly brain connectome for FAFB (female adult fly brain) – last year in preprint form, today as an immersive feature in Nature.
140,000 neurons, over 50 million synapses – organised into over 8,000 cell types. (VNC not included.)
https://www.nature.com/immersive/d42859-024-00053-4/index.html
The whole connectome: Dorkenwald et al. 2024 (Seung, Murthy) https://www.nature.com/articles/s41586-024-07558-y
Cell types: Schlegel et al. 2024 (Jefferis) https://www.nature.com/articles/s41586-024-07686-5 by @uni_matrix
Network statistics: Lin et al. 2024 (Murthy) https://www.nature.com/articles/s41586-024-07968-y
Visual system: Garner et al. 2024 (Wernet, Kim) https://www.nature.com/articles/s41586-024-07967-z and Matsliah et al. (Murthy, Seung) https://www.nature.com/articles/s41586-024-07981-1
Seung also put out a solo paper on predicting visual function from the connectome: https://www.nature.com/articles/s41586-024-07953-5
Control of halting in walking: Sapkal et al. 2024 (Bidaye) https://www.nature.com/articles/s41586-024-07854-7
FAFB imaged by @davi 's group back in 2018: https://www.cell.com/cell/fulltext/S0092-8674(18)30787-6
The FlyWire connectome: neuronal wiring diagram of a complete fly brain
Artificial intelligence and human expertise meet to generate a map of all the connections in the fly brain. The resource is already being used by experimentalists and theoreticians to further our understanding of neural circuits in the fly and beyond.