#Bacteria have a
#DNAdamage response to aberrant
#DNAmethylation, but how do they sense this damage? @Aditya_Kamat96 @Tung_BK_Le @dnarepairlab &co describe a TF that regulates a methylation-specific DNA damage response in
#Caulobacter #PLOSBiology https://plos.io/3v7pdkD 

Widespread prevalence of a methylation-dependent switch to activate an essential DNA damage response in bacteria
DNA methylation plays central roles in diverse cellular processes, ranging from error-correction during replication to regulation of bacterial defense mechanisms. Nevertheless, certain aberrant methylation modifications can have lethal consequences. The mechanisms by which bacteria detect and respond to such damage remain incompletely understood. Here, we discover a highly conserved but previously uncharacterized transcription factor (Cada2), which orchestrates a methylation-dependent adaptive response in Caulobacter. This response operates independently of the SOS response, governs the expression of genes crucial for direct repair, and is essential for surviving methylation-induced damage. Our molecular investigation of Cada2 reveals a cysteine methylation-dependent posttranslational modification (PTM) and mode of action distinct from its Escherichia coli counterpart, a trait conserved across all bacteria harboring a Cada2-like homolog instead. Extending across the bacterial kingdom, our findings support the notion of divergence and coevolution of adaptive response transcription factors and their corresponding sequence-specific DNA motifs. Despite this diversity, the ubiquitous prevalence of adaptive response regulators underscores the significance of a transcriptional switch, mediated by methylation PTM, in driving a specific and essential bacterial DNA damage response.

A new Achilles heel of the bacterial cell wall
The bacterial cell wall must be constantly remodeled in order to grow and divide. This involves the close coordination of lytic enzymes and peptidoglycan synthesis. In their study published in Nature Communications, researchers led by Martin Thanbichler have now found that a central regulator can control completely different classes of autolysins. Since many antibiotics target the bacterial cell wall, these findings may contribute to the development of new therapeutic strategies against bacterial infections.
Synchronized Swarmers and Sticky Stalks: #Caulobacter crescentus as a Model for Bacterial Cell Biology | Journal of Bacteriology
https://journals.asm.org/doi/10.1128/jb.00384-22
@goleylab on twitter:
Gorgeous image courtesy of Mabel Jones, 1905. Caulobacter-like rosettes from a bug isolated from Chicago tap water. One of my faves.
RT of @[email protected]
In this minireview, @jordanmbarrows and @goleylab explore the development and utility of Caulobacter crescentus as a model organism. #Caulobacter #BacterialCellBiology #ModelOrganisms
journals.asm.org/doi/10.1128/jb…
@ASMicrobiology
#microbiology #caulobacter
yes, this!
"Everything you ever wanted to know about #Caulobacter and its development as a model for bacterial cell and developmental biology."
Kudos to @[email protected] for this comprehensive introduction to our fave model system! says @goleylab
https://journals.asm.org/doi/10.1128/jb.00384-22
#microbiology
Phase Separation of Scaffold Protein Regulates Microbial Asymmetric Cell Division
http://microbiologycommunity.nature.com/posts/phase-separation-of-scaffold-protein-regulates-microbial-asymmetric-cell-division
Phase separation modulates the assembly and dynamics of a polarity-related scaffold-signaling hub https://www.nature.com/articles/s41467-022-35000-2
As a well-established model to study bacterial asymmetric cell division, #Caulobacter crescentus produces a motile swarmer cell and a sessile stalked cell during each cell cycle.

Phase Separation of Scaffold Protein Regulates Microbial Asymmetric Cell Division
eDNA-stimulated cell dispersion from #Caulobacter crescentus #biofilms upon oxygen limitation is dependent on a #toxin-antitoxin system
https://elifesciences.org/articles/80808

eDNA-stimulated cell dispersion from Caulobacter crescentus biofilms upon oxygen limitation is dependent on a toxin-antitoxin system
DNA life preservers.
#Caulobacter produces
#prophage-like gene transfer agent that mediates survival in DNA damaging conditions by providing genomic DNA from donor cells for
#recombination-based repair in recipients @Tung_BK_Le @michael_laub8
#PLOSBiology https://plos.io/3E0EyVR
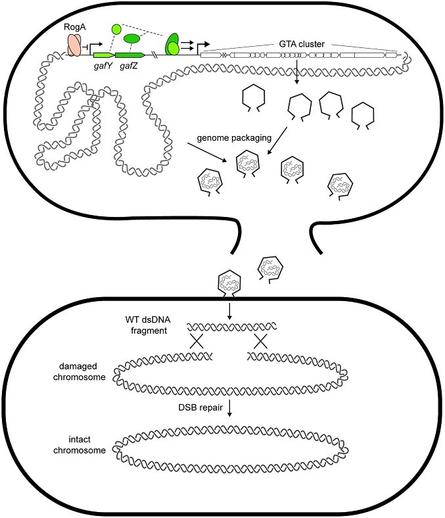
Prophage-like gene transfer agents promote Caulobacter crescentus survival and DNA repair during stationary phase
The model organism Caulobacter crescentus is found to produce a prophage-like gene transfer agent that mediates survival to stationary phase and DNA damaging conditions by providing genomic DNA from donor cells for recombination-based repair in recipient cells.
Cool work from the Brun lab - an #amyloid fiber holds the #Caulobacter super glue holdfast to the cell pole
https://doi.org/10.1128/jb.00273-22