| github | https://github.com/omaclaren |
| google scholar | https://tinyurl.com/ojmscholar |
| abandoned website/blog | https://omaclaren.com/ |
| pronouns | he/him |
Oliver Maclaren
- 506 Followers
- 557 Following
- 164 Posts
Profile-Wise Analysis: A profile likelihood-based workflow for identifiability analysis, estimation, and prediction with mechanistic mathematical models
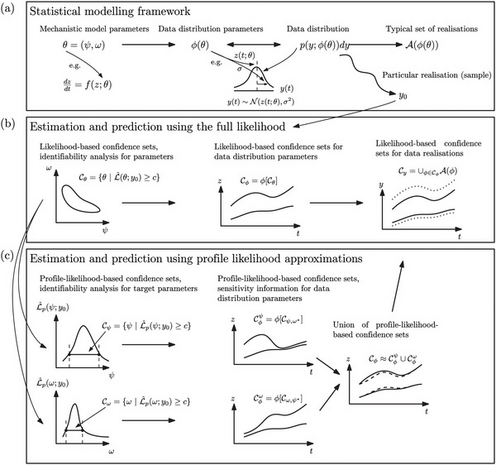
Author summary Parameter estimation and model prediction are essential steps when mathematical models are used to provide biological insight or to make practical predictions about future scenarios. We present an efficient, unified workflow that addresses parameter identifiability, parameter estimation and model prediction from a likelihood-based frequentist perspective. Our workflow, called Profile-Wise Analysis (PWA), involves constructing ‘profile-wise’ predictions that propagate profile-likelihood-based confidence sets for model parameters to predictions, explicitly isolating how different parameter combinations affect model predictions. Combining profile-wise prediction confidence sets gives an overall prediction confidence set that efficiently approximates the full likelihood-based prediction confidence set. Three case studies, focusing on canonical mathematical models used in biology and ecology, illustrate various aspects of the workflow for commonly-encountered ODE-based mechanistic models with both Gaussian and non-Gaussian measurement error.