| Website | https://nextstrain.org |
| Bluesky | https://bsky.app/profile/nextstrain.org |
| https://twitter.com/nextstrain | |
| Github | https://github.com/nextstrain |
Nextstrain
- 2K Followers
- 0 Following
- 62 Posts
To facilitate discovery of datasets and guide users to the most suitable dataset, Nextclade quickly compares kmers from the input sequence to an index and highlights good matches.
The CLI has a sub-command `sort` that will split input sequences into subsets that match different reference sequences.
[4/n]
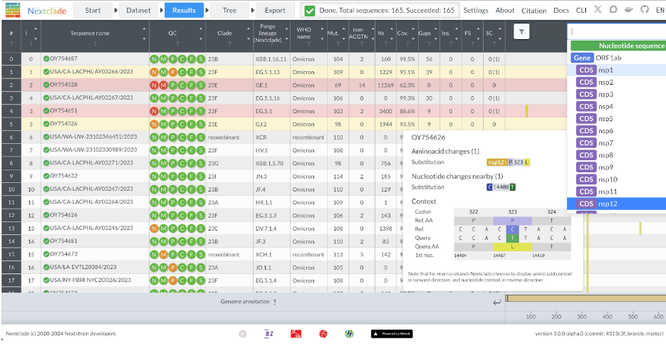
Instead of just attaching sequences to the reference tree, Nextclade now resolves phylogenetic structure in the submitted sequences. Makes the tree view much more informative.
Caveat: it’s a greedy tree builder that’s not a substitute for proper phylogenetics
[3/n]
With the support for splicing and slippage, we can now provide a dataset that annotates all mature proteins of SARS-CoV-2.
https://master.clades.nextstrain.org/?dataset-name=nextstrain/sars-cov-2/wuhan-hu-1/proteins
Going forward, this enables datasets for HIV or Hepatitis B virus.
[2/n]
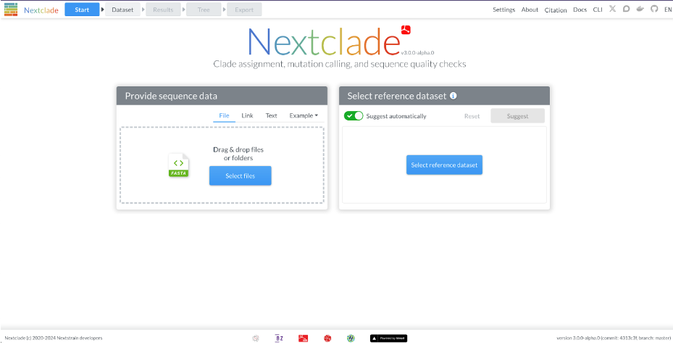
After many months of work, we are releasing Nextclade version 3 with major new features and changes:
- Splicing, slippage, and circular genomes
- Local tree-building
- Auto-suggestion of suitable datasets
- Align more divergent sequences
- Community datasets
clades.nextstrain.org
[1/n]
Additionally, we've updated the Nextclade datasets to appropriately call clade 23G, 23H, and 23I viruses at clades.nextstrain.org.
If you use #Nextclade in the browser, make sure to reload the tab from time to time so that you get the most recent datasets.
8/9
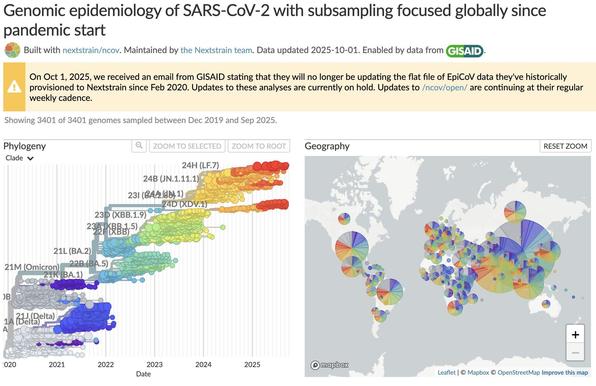
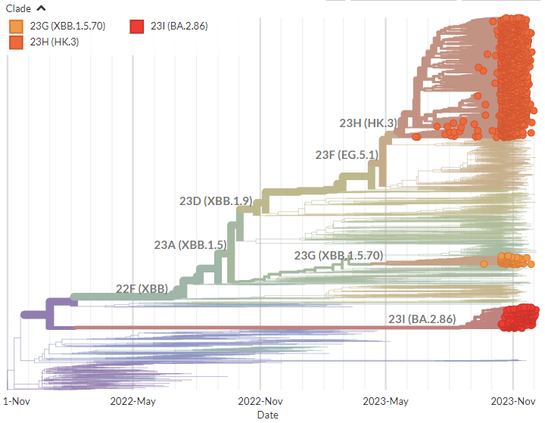
You can take a closer look at 23G, 23H, and 23I in the 2-month Nextstrain build below:
6/9
23I (BA.2.86) sublineage JN.1 (sometimes referred to as 'Pirola') has extra RBD mutation L455S & shows a further increase in growth. It likely emerged in Europe, where it is most common, and makes up around half of all 23I sequences.
5/9
23I (BA.2.86 - *not* an XBB descendant) represents a significant evolutionary jump, comparable to the emergence of Omicron, with over 30 spike mutations more than its BA.2 ancestor.
It likely emerged in Southern Africa in mid-2023, and has grown rapidly worldwide.
4/9
23H (HK.3) is the largest sublineage of 23F (EG.5.1), making up more than half of 23F sequences. With the addition of L455F on top of the F456L from 23F also giving it the FLip genotype.
It emerged in mid-2023, likely in China, and is particularly common in Asia.
3/9