Great new addition to the #preLights5thBirthday #preList!
Samantha Seah, Navin B Ramakrishna & Hui Ting Zhang #astar_gis highlight a #preprint from the #FergusonSmith Lab.
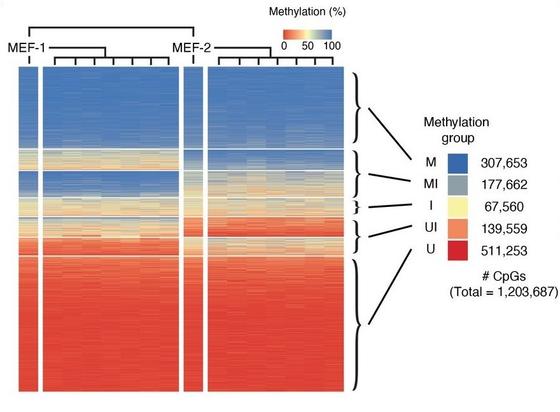
Probabilistic inheritance of intermediate DNA #methylation sites across somatic cell division