Joel Lüthi
- 83 Followers
- 126 Following
- 10 Posts
This year, the hackathon will happen in the context of the Zarr V3 transition and how we best manage this. Main topics will include OME-Zarr in Java, OME-Zarr in the Python ecosystem, OME-Zarr workflows & more. Learn more about the program here: https://biovisioncenter.notion.site/OME-NGFF-Workflows-Hackathon-2024-dde32a032adf49b4a53b4b014586b678
We have structured the hackathon to have 3 core days from Tuesday - Thursday, with the option to attend a (TBD) Monday afternoon program & an extra hacking day for Friday for super motivated people.
(2/3)
I'm very excited to announce the #OME_NGFF_workflows_hackathon for November 18th - 22nd at the BioVisionCenter in Zurich, Switzerland.
Sign up here: https://ema.uzh.ch/en/register/ome-ngff-workflows-hackathon-2024.html
This event is a follow-up to last year's successful hackathon, where 45 researchers and developers pushed the boundaries of next generation bioimage formats, processing & workflows. Learn more about last year's event here: https://forum.image.sc/t/outcomes-of-the-next-generation-bioimage-analysis-workflows-hackathon/88733
(1/3)
(4/5)
(2/5)
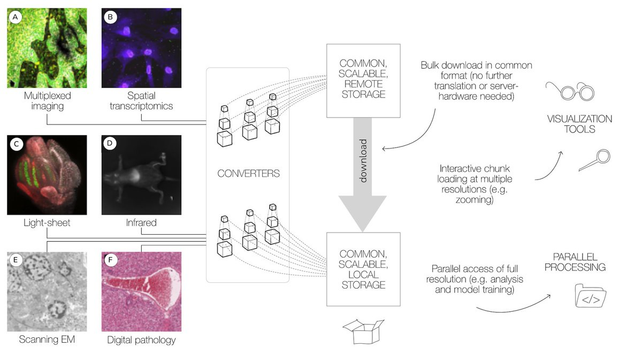
OME-NGFF: a next-generation file format for expanding bioimaging data-access strategies - Nature Methods
OME’s next-generation file format (OME-NGFF) provides a cloud-native complement to OME-TIFF and HDF5 for storing and accessing bioimaging data at scale and works toward the goal of findable, accessible, interoperable and reusable bioimaging data.
Super happy that the preprint for the #OMEZarr paper is out now! If you want to know where bioimage analysis is going: I think this will be the direction! Standardization & open source for the win!
Find the #bioRxiv preprint here: https://www.biorxiv.org/content/10.1101/2023.02.17.528834v1
And Josh's main thread about it here: https://fediscience.org/@joshmoore/109904124384328635
Thread (1/5)
Latest report from the #OMENGFF #Community describing #OMEZARR, a #CloudOptimized #BioImaging #DataFormat, is out now on #bioRxiv!
If you are having trouble with big, slow, or otherwise unwieldy imaging data, reach out or take a look for a #FAIR-er option.