Bram van Dijk
- 248 Followers
- 145 Following
- 116 Posts
Woah, the list of MGEs keeps growing! 🤯
Tycheposons! But is it pronounced tie-key, tie-chi, or jif?
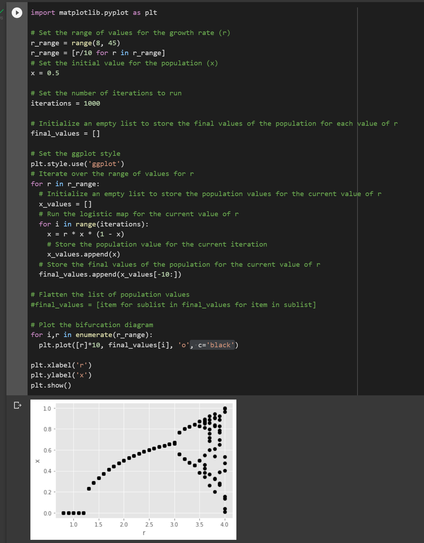
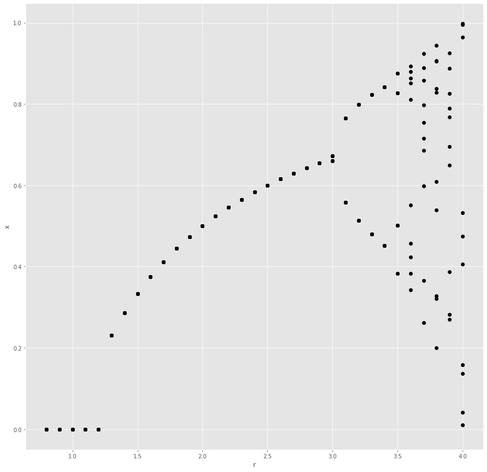
ChatGPT: "Write is a piece of code that will generate a bifurcation diagram of the logistic map in Python. The value for r run from 0.8 to 4.5. Plot the last 10 values in the iteration as dots using matplotlib with the ggplot theme."
Result below.
Vertical and horizontal gene transfer tradeoffs direct plasmid fitness - PubMed
Plasmid fitness is directed by two orthogonal processes-vertical transfer through cell division and horizontal transfer through conjugation. When considered individually, improvements in either mode of transfer can promote how well a plasmid spreads and persists. Together, however, the metabolic cos …
Evolutionary rescue of resistant mutants is governed by a balance between radial expansion and selection in compact populations - Nature Communications
Antibiotic and anti-cancer therapy are challenged by mutation-mediated treatment resistance despite many mutations being maladaptive. Here, the authors introduce a system that shows how the probability of the long-term persistence of drug-resistant mutant lineages can be increased in dense microbial populations by acquiring multiple mutations.
Big cheers to all the coauthors, #DarshakBhatt, #ToosDaemen and #FranjoWeissing
Link: https://doi.org/10.1371/journal.pcbi.1010076
Modelling the spatial dynamics of oncolytic virotherapy in the presence of virus-resistant tumour cells
Author summary Oncolytic virotherapy is a promising form of cancer treatment that uses viruses to target, infect and kill cancer cells. Unfortunately, this form of therapy is often not effective, due to the occurrence of virus-resistant tumor cells. As it is challenging to assess the emergence and spread of resistance experimentally or in (pre)clinical studies, we designed a model that allows to study the spatial dynamics of virus-sensitive and virus-resistant tumor cells in various scenarios, and to predict the efficacy of virotherapy. By analysing the model systematically, we demonstrate the importance of 2D and 3D spatial interactions, the effects of viral properties (such as replication rate and range of infection), the properties of virus-resistant cancer cells (such as the cost of resistance), and the sensitivity of healthy (non-tumor) cells towards viral infection. Our goal is to provide a sound conceptual understanding of the mechanisms underlying therapeutic failure, which eventually may lead to the discovery of strategies that improve therapeutic efficacy. We therefore provide the reader with a graphical and a terminal interface of our model (executable on a local computer), allowing practitioners to reflect on their intuition regarding the complex yet fascinating dynamics of oncolytic virotherapy.
Environmental and ecological controls of the spatial distribution of microbial populations in aggregates
https://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1010807
Environmental and ecological controls of the spatial distribution of microbial populations in aggregates
Author summary Microbial communities are assembled by the interactions between microorganisms and the local environment. To fully understand and control the formation of microbial aggregates, we need to unravel the principles of both cell-cell, cell-environment and cell-space interactions. Until now, most studies have focused predominantly on single interactions between two microbes. However, microbial ecology is more complex than that, and multiple ecological interactions contribute to microbial community assembly. The identification of distinct spatial distributions of bacteria is a first step towards the understanding the underlying biological mechanisms that govern aggregate formation. Here, we show that it is possible to evaluate the influence of multiple ecological interactions and the environment on microbial community assembly through mathematical modelling. We have been able to distinguish interspecific segregation of communities in competition, and layered distribution in commensalism. When we considered more than one ecological interaction between populations, the resultant spatial distribution was identified as the one controlled by the most limiting substrate. Additionally, we defined a theoretical modulus that able us to predict the most probable spatial distribution under specific environmental conditions.