A spoonful of sugar #18 🍬
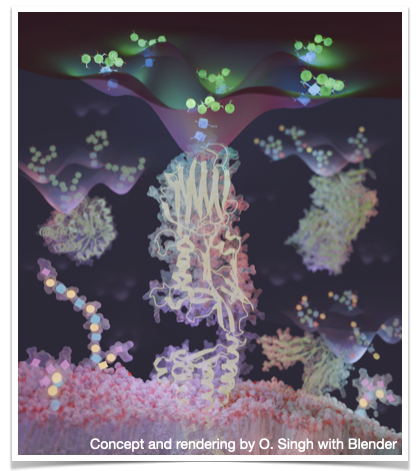
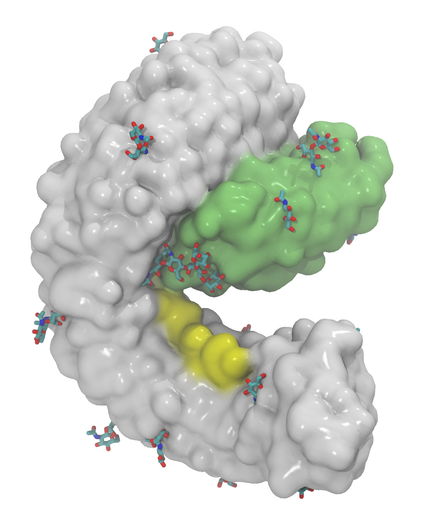
Plant receptor kinases translate and pass on chemical messages as part of many key biological processes. A new fabulous cryo-EM structure of the active complex between the kinase MIK2, the immunomodulatory peptide SCOOP and the co-receptor BAK1 shows how its binding hinges on N-glycosylation (https://doi.org/10.1038/s41477-024-01841-6)
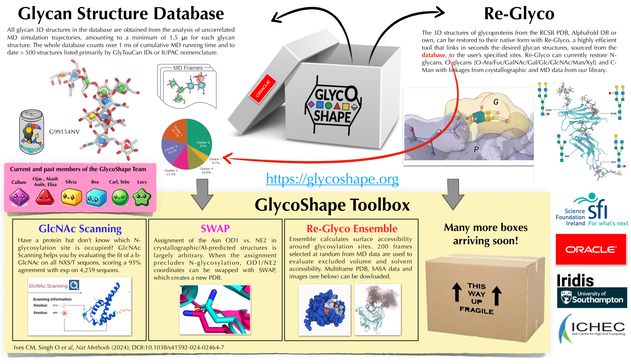
You can rebuilt the fully glycosylated structure of the complex with https://glycoshape.org with some fancy-plant N-glycans 
 BIG MASSIVE THANK YOU 👏 to all the beta testers around the world. Happy glycosylating!
BIG MASSIVE THANK YOU 👏 to all the beta testers around the world. Happy glycosylating!