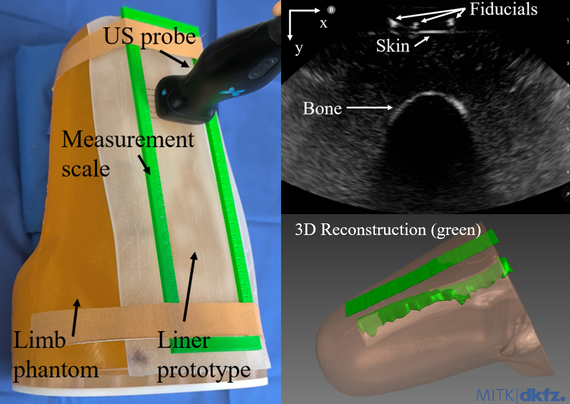
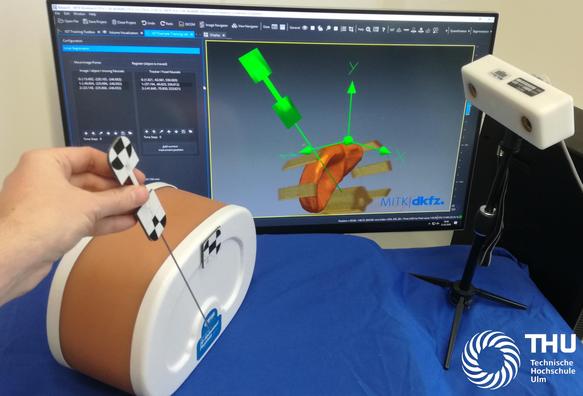
Prototypes for medical navigation often require complex hardware and software setups. Students at THU have developed a Java-based example that can be easily compiled. It is built using Gradle and can be connected to other tools such as MITK-IGT or PLUS via OpenIGTLink. At BVM they demonstrated a use case for stroke research: A catheter in a brain vessel dummy was localized using AI in simulated fluoroscopy images and navigated to a target.
| Web | https://www.thu.de/franz |