[5/13]
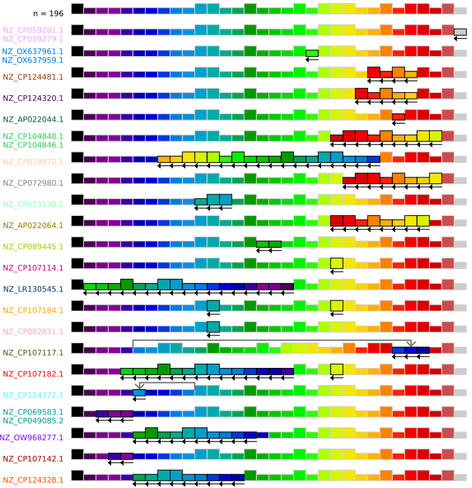
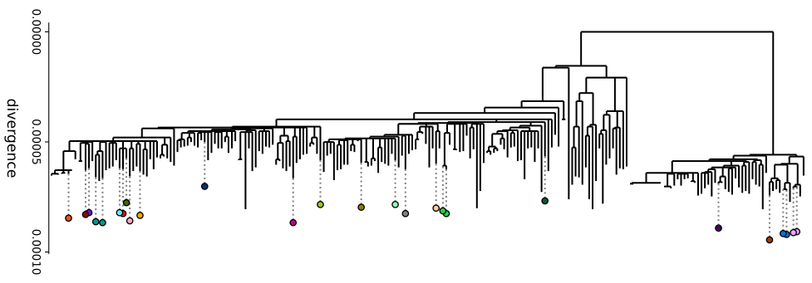
Core-genome synteny, i.e. the order of core genes, has been shown to be strongly conserved across large evolutionary distances. But what happens on short timescales? Using the graph we can survey all possible changes of synteny in the dataset. Out of 222 isolates, we find only 26 with any change in synteny. Most of these changes are inversions, often around the origin or terminus of replication.
[6/13]
This suggests that changes in synteny are relatively frequent, occurring at a rough rate of one every ~250 clonally-inherited mutations on the (filtered) core-genome. However, the fact that synteny is largely conserved across big evolutionary distances, and the fact that many of these changes happen on terminal branches of the tree, indicate that these changes are likely removed by purifying selection.
[7/13]
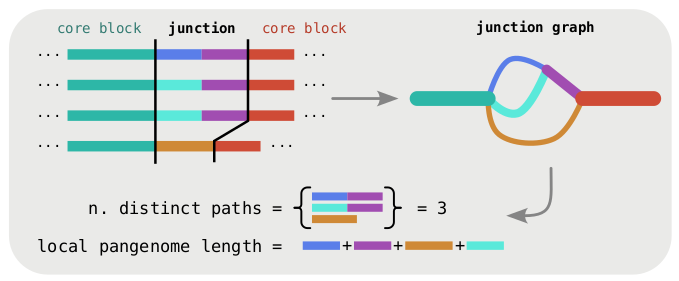
Next we investigate the structural diversity in the accessory genome. The fact that the order of core-genes is mostly conserved provides a well-defined frame of reference in which to study accessory variation. We look at the local graph between two adjacent core blocks, that we call a junction graph. In this graph the diversity can be quantified in terms of number of distinct paths and total accessory sequence content.
[8/13]
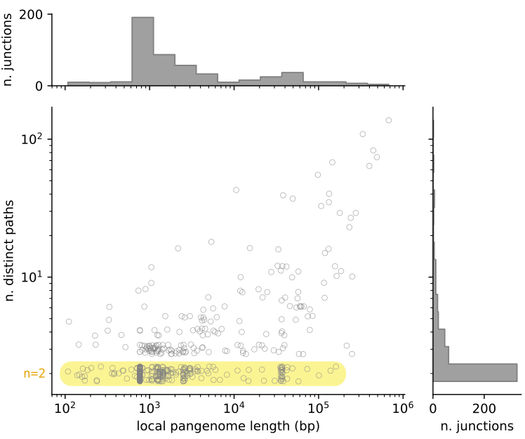
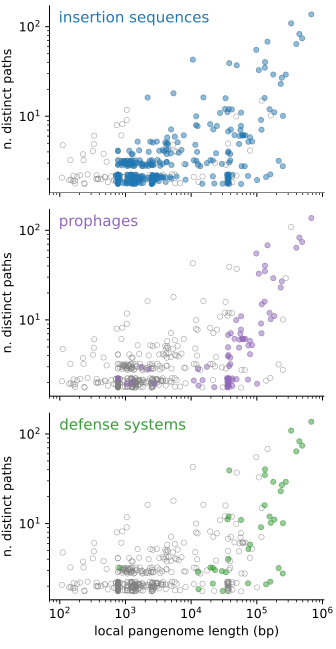
If we scatter-plot these two quantities for all of the 519 junctions in the dataset, we find that the majority of junctions are binary, i.e. they contain only two possible distinct paths, of which one is often empty. Their length distribution is bimodal, with a peak around 1kbp and another around 30-40 kbp. On the other end of the spectrum we find hotspots, regions with tens to hundreds of different distinct paths, and a total accessory genome content of tens to hundreds of kbp in length.