This weeks #ChemSciPicks is a fantastic Edge article from Charlotte Deane et al., (University of Oxford).
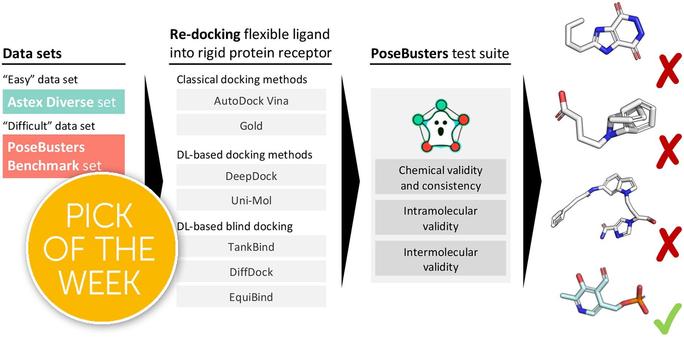
This Edge article reports PoseBusters, a Python package that performs a series of standard quality checks using the well-established cheminformatics toolkit RDKit.
You can read the work for free here:
https://doi.org/10.1039/D3SC04185A
(This was the paper that I was excited to share and I am so glad that the rest of the Chemical Science editorial team agreed with me!)

 )
)