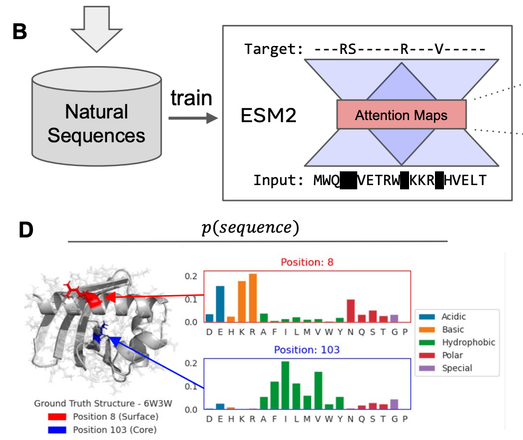
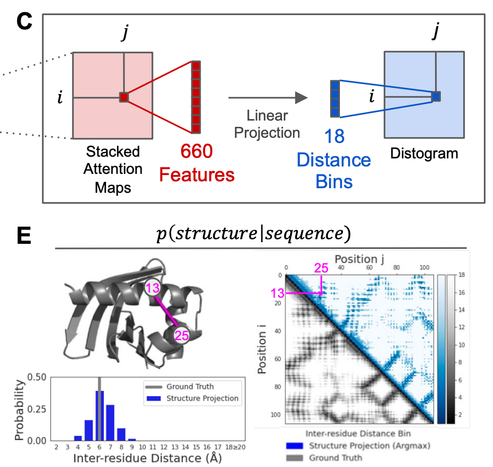
Given the observation that attention maps in the LMs correspond to contacts. One can train a linear projection from the attention maps to a distogram, allowing the modeling of P(structure | sequence). (2/5)
Papers showing LMs learn contacts:
https://arxiv.org/abs/2006.15222
https://www.biorxiv.org/content/10.1101/2020.12.15.422761v1
https://www.biorxiv.org/content/10.1101/2020.12.21.423882v2
BERTology Meets Biology: Interpreting Attention in Protein Language Models
Transformer architectures have proven to learn useful representations for protein classification and generation tasks. However, these representations present challenges in interpretability. In this work, we demonstrate a set of methods for analyzing protein Transformer models through the lens of attention. We show that attention: (1) captures the folding structure of proteins, connecting amino acids that are far apart in the underlying sequence, but spatially close in the three-dimensional structure, (2) targets binding sites, a key functional component of proteins, and (3) focuses on progressively more complex biophysical properties with increasing layer depth. We find this behavior to be consistent across three Transformer architectures (BERT, ALBERT, XLNet) and two distinct protein datasets. We also present a three-dimensional visualization of the interaction between attention and protein structure. Code for visualization and analysis is available at https://github.com/salesforce/provis.
@neuropunk
It's good for the structure prediction task, you want the model to be robust and recognize the bare minimum signal from the input sequence. But not good for the design task.
Let's say you have a suboptimal sequence that only partly encodes the desired structure. If your model is "too good", it will fill in the rest of the structure. (1/2)
@neuropunk
So now, during design, as soon as you get the desired structure, there is no longer any signal to update your sequence.
In this case, we wanted the structure to be fully encoded in the LM's contacts, and to avoid a situation where a more complex structure module starts hallucinating or improvising. (2/2)