Citations in scientific presentations should be readable and easy to follow for the audience even at the back of the room.
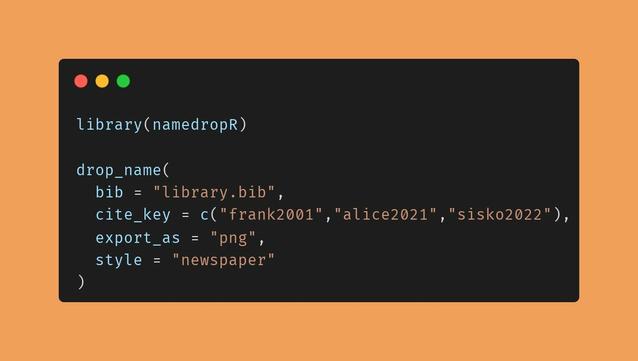
I wrote the tiny #rstats package {namedropR} to make this easy to achieve for speakers.
A quick 'How to' is in the thread below.