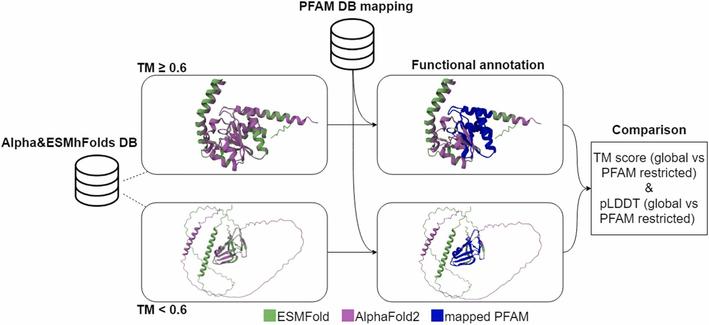
🧬 How do AlphaFold2 and ESMFold stack up when it comes to functional annotation?
🔗 AlphaFold2 and ESMFold: A large-scale pairwise model comparison of human enzymes upon Pfam functional annotation. Computational and Structural Biotechnology Journal, DOI: https://doi.org/10.1016/j.csbj.2025.01.008
📚 CSBJ: https://www.csbj.org/
#AlphaFold2 #ESMFold #AIinScience #ProteinStructure #Enzymes #Pfam #UniProt #AlphaFold #HumanProteome #FunctionalAnnotation #AIinBiology #ComputationalBiology #FunctionalGenomics