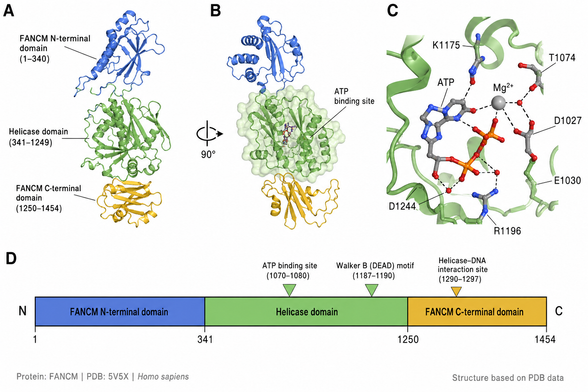
FANCM (PDB: 5V5X) — DEAD-box ATPase geometry (K1175/Mg²⁺) drives replication fork reversal at interstrand crosslinks, not classical duplex unwinding. Helicase–DNA interface: residues 1290–1297.
Fork reversal fidelity declines with age. FANCM's structural landscape is an underexplored target in replication stress and genomic aging.
#FANCM #FanconiAnemia #StructuralBiology #DNARepair #ReplicationStress #LongevityBiology #Biogerontology