I'm pleased to announce that antiSMASH can now predict if an NRPS BGC produces a metallophore!
Check out our new preprint for all the details:
https://www.biorxiv.org/content/10.1101/2022.12.14.519525v1
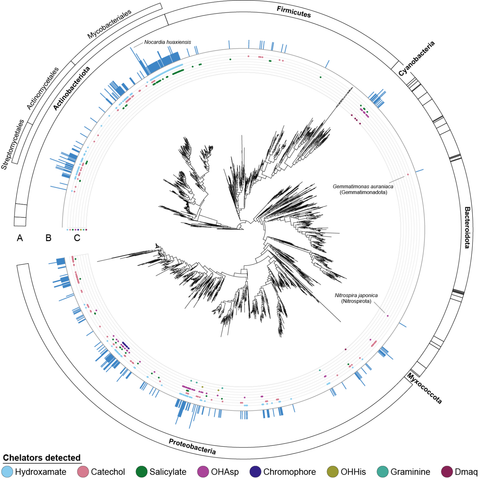
I tested the new algorithm on 15k bacterial genomes. NRP metallophores make up an astounding 25% of all NRPS BGCs! This thread will have some more highlights.
Test it out on the antiSMASH website and let me know what you think!
https://antismash.secondarymetabolites.org/#!/start
(You may have to clear your cache for "Enable antiSMASH beta" to appear)