i've been working nonstop for so many months on the same problem that when i finally think i've solved it, it kind of triggers a state of shock, more than anything else... i think i may actually be in shock!
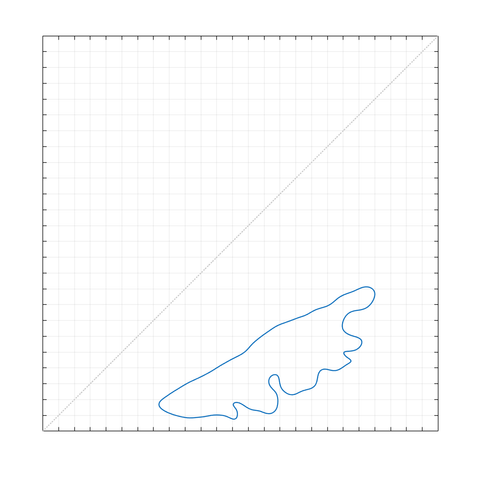
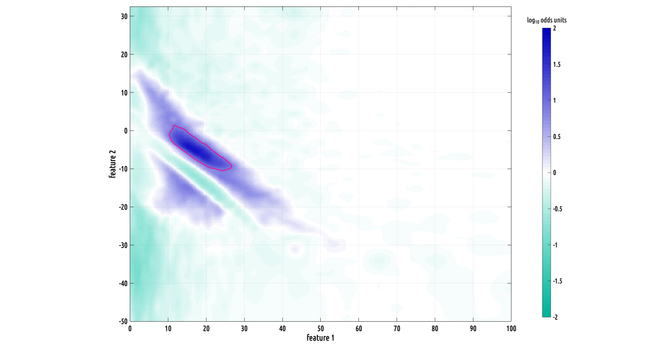
here's a preview of what i've found using this new technique... happily, this is grant work, so i will be able to publish more on these results soon! as you can see, this outline indicates the kind of shape that would be difficult to characterize using a parametric fitting method.
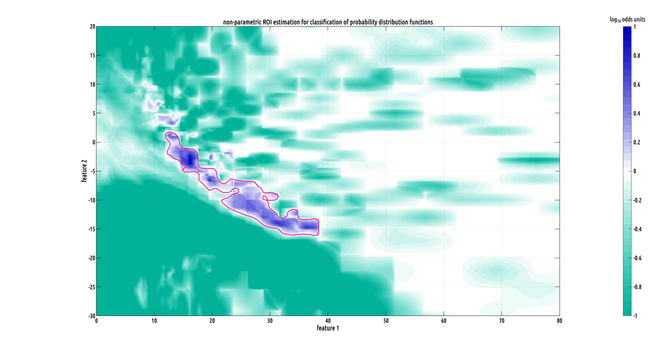
Still working on the writeup but I did make a slightly better plot :) the problem we had was that our data has the form of probability distribution functions over a 2D feature space from two different classes, and they have a lot of variation from subject to subject within a class, making binary classification challenging.
what contiguous regions in our 2D feature space are significantly and consistently shifted from each other between classes? i formulated a nonparametric test to answer this.
there are about 15,000 hours of electrographic recordings over roughly a 3 week period using 30 subjects going into these distribution estimates. our waveform-shape-analytic techniques extract about 100K events per hour. that way, even probabilities of relatively rare events, like 1 in 100,000, can be accurately combined into a (relatively) smooth picture such as this one.
the precision of these distribution estimates allows us to lower the effective noisefloor of the recordings. in other words, these density-functional ROIs enable us to pull highly specific signals out, that would otherwise be buried beneath the noisefloor.
that's the good news. the bad news is that the pathology we're trying to detect is sparsely represented across electrodes. so, you could have less than 30% of your subjects presenting on one electrode, but without it, your sensitivity is shot
this makes combining information across electrodes particularly interesting! happily, if all of your signals are close to 100% specific, you don't have to work too hard to combine a sparse set of them together. (e.g. the max, aka the probabilistic/fuzzy OR operator.) or, you can also use a personalized approach, by inferring which electrodes to base your prediction on for each subject, based on their features.
so, it's not something we are too worried about, although much larger datasets are required for us to better validate the results we are seeing from this initial study. hopefully, we'll be able to publish our grant report, or some aspect of it, soon...
(in case you're wondering, simply assembling the ROI features from each electrode into a standard feature matrix wouldn't be easy with this dataset because of outliers that were not homogeneous across recording channels, but that would normally be an option)
assuming that we get funding for clinical trials & get approval, we envision delivering ROIs like these to epileptologists that are unable to currently make a diagnosis with only one hour of EEG data, requiring epilepsy monitoring unit (EMU) visits for longer (3-7 day) electrographic recordings. from what our advisors are telling us, this is a big bottleneck in diagnosing epilepsy rapidly, because EMUs are backlogged and it can be a 6 month wait (or more). also, not every area has an EMU nearby.
another big need is biomarkers of epileptogenesis, prior to epilepsy onset. that's what I've been working on for the last couple of years... digital biomarkers for prediction of post-traumatic epilepsy in traumatic brain injury. we have some really interesting preclinical results, but that is an even bigger challenge than rapid epilepsy diagnosis. so, if all goes well, we hope to launch a diagnostic test first, and then a predictive test second.
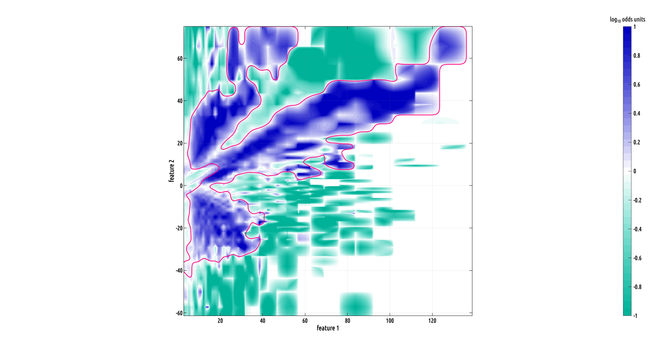
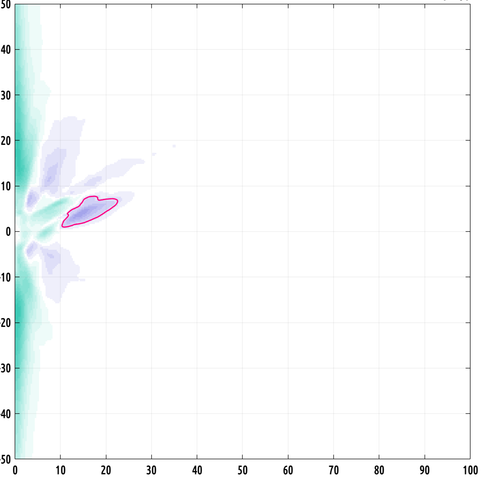
here's a really cool looking region of interest (ROI). this one is big!
the formula i came up with works in any dimension. of course, explicit density estimation rapidly breaks down with the curse of dimensionality. so, we probably can't extend this to distributions on high dimensional domains... but I can't wait to try this in 3D when I have more time :)
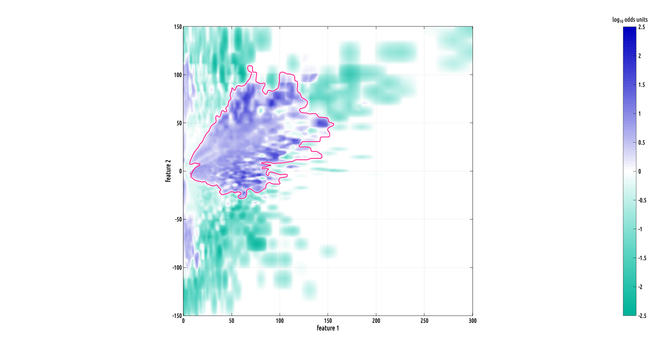
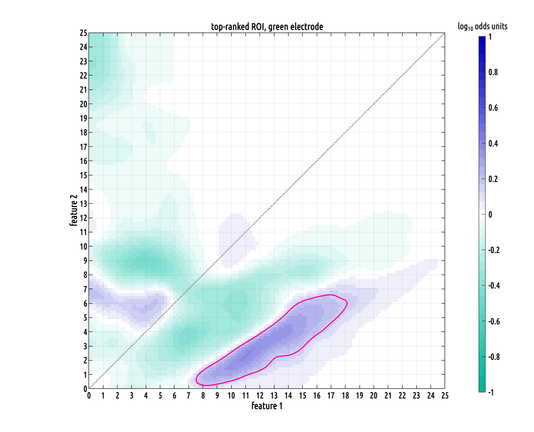
each subject contributes one 2D distribution to the set, so intra-class variability between proximal points in 2D can be large. one of the cool things about this nonparametric ROI test is that it has built-in robustness parameters. this means that it is capable of finding meaningful ROIs even in the presence of outliers and low sensitivity signals among the samples within groups. but, these parameters also make it difficult to visualize what data the ROIs are based on. here's a much larger area
what's really interesting to me is noticing patterns in the geometry of these novel EEG biomarkers of epilepsy. it seems to me that each consistent geometric pattern in the differences of these probability distribution between classes could indicate some type of physiologically-relevant phenotype, especially when it is consistent across different studies / recordings / epilepsy types.
as someone new to the field, there is, ironically, a risk associated with being too innovative. namely, not everyone will believe your results :) that's probably been our biggest barrier to date: "the dataset is too small", or "we've seen a lot of claims to this effect, that have all proven fraudulent. how were you able to succeed?". after several years of these kinds of reactions, getting our first grant was a really big deal for us. but now that the money has run out, next steps are unclear
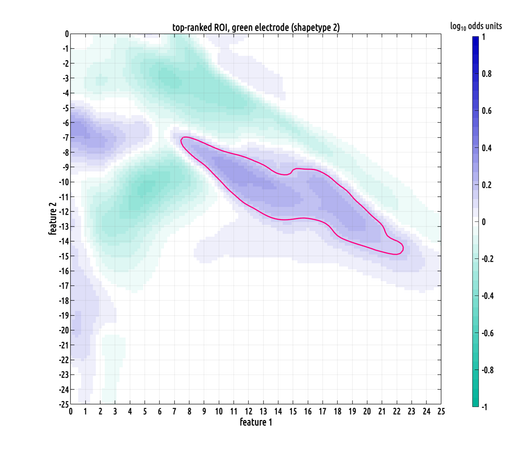
i'm still fine-tuning the design of the test. there are a lot of adjustment you can make, but in the interest of minimizing parameter number, four will hopefully be enough! this is the most specific ROI we've found for post-traumatic epilepsy in this dataset.
in essence, this formula allows us to infer topologic relations from topographic ones
i apologize if this thread is too long, it's because i've been working on this problem for so long, that to see such beautiful pictures alongside the empirical performance evaluation results i'm seeing is such a huge validation of the 6+ years i've put into this project.
#phew