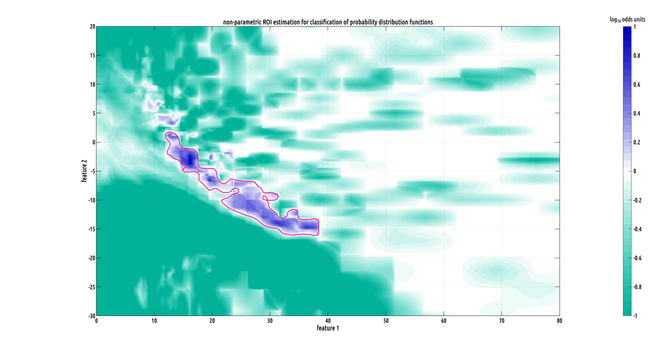
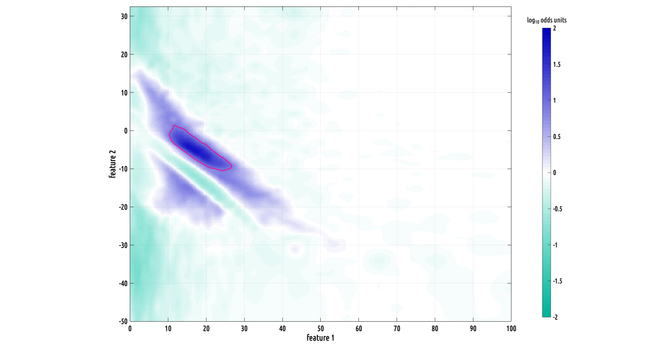
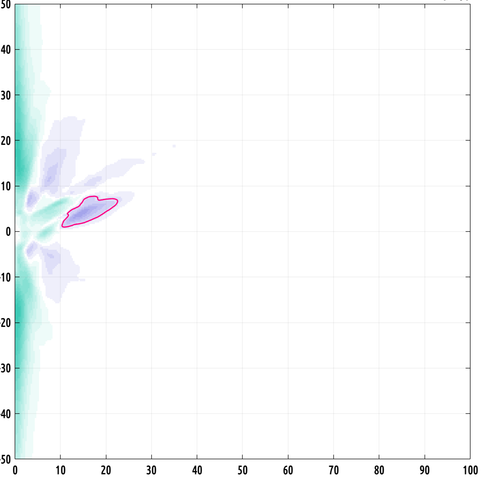
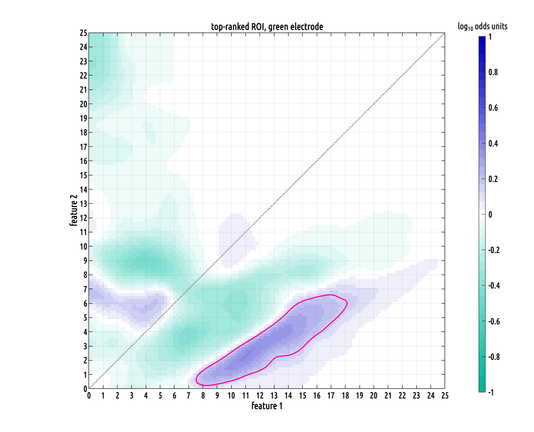
Still working on the writeup but I did make a slightly better plot :) the problem we had was that our data has the form of probability distribution functions over a 2D feature space from two different classes, and they have a lot of variation from subject to subject within a class, making binary classification challenging.
what contiguous regions in our 2D feature space are significantly and consistently shifted from each other between classes? i formulated a nonparametric test to answer this.
the precision of these distribution estimates allows us to lower the effective noisefloor of the recordings. in other words, these density-functional ROIs enable us to pull highly specific signals out, that would otherwise be buried beneath the noisefloor.
that's the good news. the bad news is that the pathology we're trying to detect is sparsely represented across electrodes. so, you could have less than 30% of your subjects presenting on one electrode, but without it, your sensitivity is shot
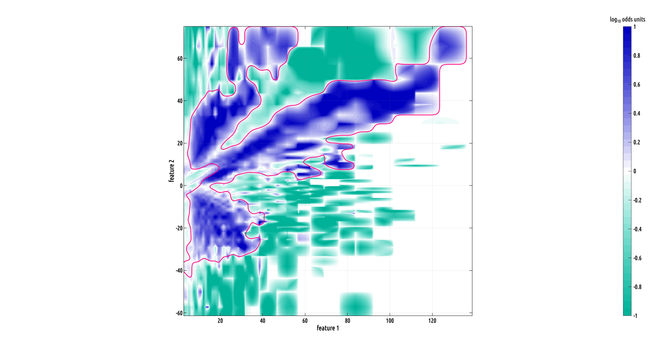
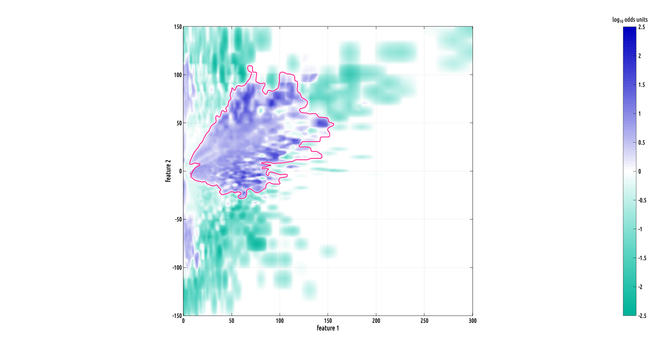
here's a really cool looking region of interest (ROI). this one is big!
the formula i came up with works in any dimension. of course, explicit density estimation rapidly breaks down with the curse of dimensionality. so, we probably can't extend this to distributions on high dimensional domains... but I can't wait to try this in 3D when I have more time :)
when i was first studying linear algebra, my Dad always tried to help me. but i was so, so far below his level, that most often i could not understand what he was trying to tell me. (i'm not mathematically gifted in the way that he was.)
the 2-D-first decision was a high-level decision that I made myself, based on 20+ years of experience. no business manager, MBA or otherwise, would do this.
that is a microcosm of this entire project: it's been R&D led. and i can't wait to see where it goes.
Oh, yeah, I mentioned my Dad and linear algebra. well, what he told me was that pretty much any proof in linear algebra can be extended from a 2-D proof. so, by working in 2-D, you're not really constraining yourself too much.
I think that the same is true for feature engineering. if your representation is extensible to N-D, but you haven't done the 2-D R&D work thoroughly before extending it, you're messing up.