RE: https://fediscience.org/@Guillawme/111534984107819771
Something really cool happened to me this year!
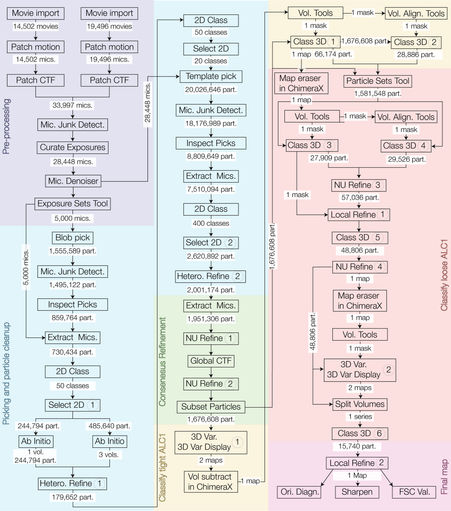
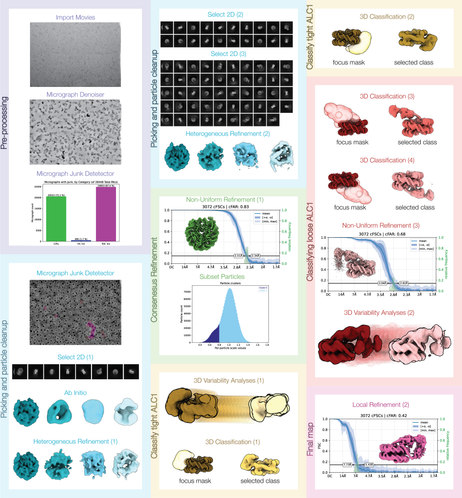
@HRBridges re-processed a #cryoEM dataset from some previous work of mine and colleagues (publicly available as EMPIAR-10739; see quoted post below for a summary of this work). She significantly improved the results we had struggled to obtain at the time of the initial analysis, and wrote it up as a case study for the CryoSPARC Guide: https://guide.cryosparc.com/processing-data/tutorials-and-case-studies/case-study-end-to-end-and-exploratory-processing-of-a-motor-bound-nucleosome-empiar-10739
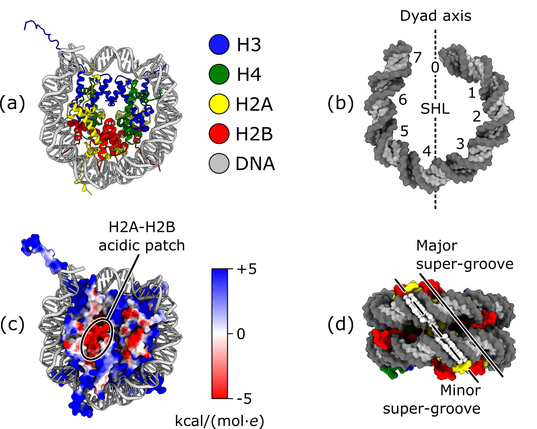
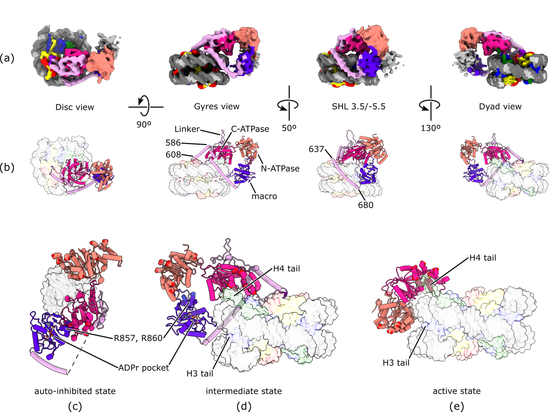
This is already super cool! Even cooler: she found a structure we had not found in this dataset. And it is a significant piece of result both to complete our understanding of ALC1 (the protein under study) and more broadly to understand one more way in which chromatin-binding factors can recognize the #nucleosome
We wrote an article about it, the preprint went online in November (now trying to get it peer-reviewed, but it might take some time; I can only work on this in my free time): https://doi.org/10.1101/2025.11.10.687450
I wrote some more about the back story here: https://www.gaullier.org/en/blog/2025/12/28/new-preprint-on-alc1
Long story short: this preprint would not have materialized, had we not both attended the CCP-EM Spring Symposium this year and discussed at the poster session.
I will write a summary thread about this new preprint in the near future (hopefully before the spring semester hits me; it's going to be busy...).