Our lab has published a couple of major papers in the past few months, which I would like to highlight in this thread!
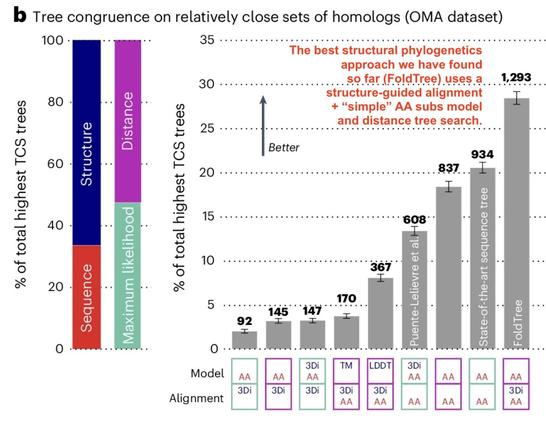
Structure-informed phylogenetic inference with FoldTree outperforms sequence-only trees on divergent proteins, and untangles RRNPPA quorum-sensing receptors across Gram-positive bacteria & phages. Led by David Moi (https://www.linkedin.com/in/david-moi) Paper: http://doi.org/10.1038/s41594-025-01649-8
Testing the “least-diverged ortholog” conjecture at scale: using >1M structures + expression across 16 animals & 20 plants, the LDO tends to retain ancestral function while the other copy specialises. Led by Irene Julca (https://www.linkedin.com/in/irene-julca-ph-genomics) Paper: http://doi.org/10.1101/gr.280166.124 Preprint: https://www.biorxiv.org/content/10.1101/2024.10.29.620890v1.full.pdf+html