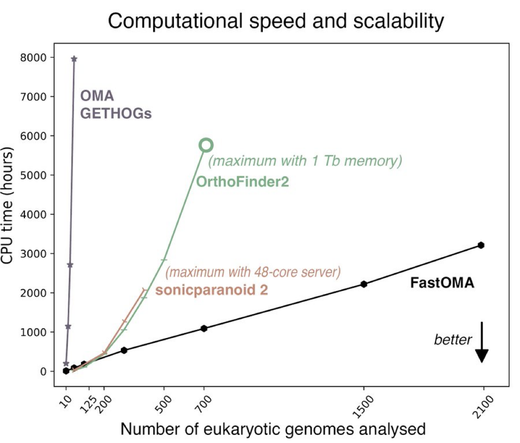
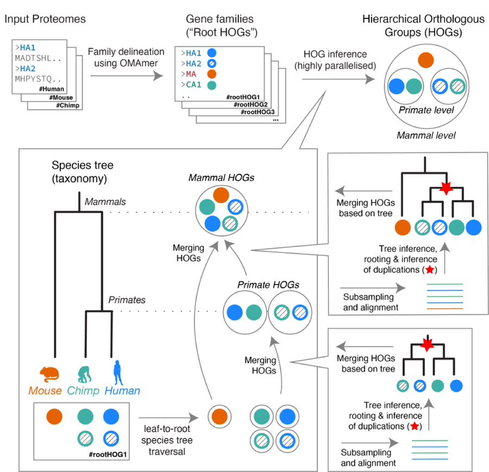
Our lab is excited to release FastOMA, a rewrite of OMA orthology algorithm that scales linearly, while maintaining OMA’s high accuracy. FastOMA can process all >2000 eukaryotic UniProt ref proteomes within 24 hours. Try it out https://github.com/DessimozLab/FastOMA
Preprint https://www.biorxiv.org/content/10.1101/2024.01.29.577392
Preprint https://www.biorxiv.org/content/10.1101/2024.01.29.577392